Sense, Antisense: C9ORF72 Makes Both Forms of RNA, Peptides
Quick Links
The puzzle of neurodegenerative disease caused by expansions in the C9ORF72 gene just grew more intricate. In four recent papers, researchers report that the repeat is transcribed and translated not only in the forward, sense direction, but also in the backward, antisense direction. The findings suggest that any therapy—such as antisense oligonucleotide treatment—may have to address both sense and antisense C9ORF72 sequences. “We have just doubled the complexity,” said John Ravits of the University of California in San Diego, senior author of one study published in the October 29 Proceedings of the National Academy of Sciences online. The other studies appeared in Acta Neuropathologica throughout October.
C9ORF72 hexanucleotide expansions are responsible for inherited forms of both frontotemporal dementia and amyotrophic lateral sclerosis (see Sep 2011 news story). After two years of study, researchers have three hypotheses for the mechanism of disease. One, expansions interrupt normal gene expression, causing a dearth of the C9ORF72 gene product, whose function remains uncertain. This is the haploinsufficiency idea. Two, expansion transcripts aggregate into RNA foci that sequester crucial RNA-binding proteins (see Jan 2013 news story). Three, translation of the repeat creates abnormal, potentially toxic peptides (see Feb 2013 news story and news story). This is the gain-of-toxic-function idea. The new studies support the latter by adding antisense RNA foci and antisense peptides to the mix. However, a fifth paper suggests that the expanded C9ORF72 gene carries methyl groups, which can silence transcription, supporting the haploinsufficiency idea.
Dual-Direction Transcription and Translation
Other repeat-expansion disorders, such as Huntington’s disease and spinocerebellar ataxia, show evidence of antisense transcription of affected genes (see Cho et al., 2005, Moseley et al., 2006, Ladd et al., 2007, Chung et al., 2011, and Wilburn et al., 2011). Based on this precedent, it was logical to look for antisense transcription in C9ORF72 disease, said Adrian Isaacs of University College London, senior author of one of the Acta Neuropathologica papers. First author Sarah Mizielinska and colleagues used fluorescence in situ hybridization (FISH) to identify both sense and antisense C9ORF72 transcripts in tissues from people who died with expansion-based ALS or FTD. Both kinds of transcript formed RNA foci in the nuclei, and occasionally in the cytoplasm, of neurons from the frontal cortex as well as from the hippocampus and cerebellum. RNA foci were less abundant in astrocytes, microglia, and oligodendrocytes. In addition, Isaacs and colleagues observed that the number of RNA foci correlated with age of disease onset. People with more foci had gotten ill at a younger age, adding to the evidence that the RNA foci play a role in the disease, said Isaacs. These data support the toxic RNA theory of C9ORF72 disease, Isaacs said, but he cautioned that other mechanisms remain possible. Ravits and colleagues also report both sense and antisense RNA foci in cortical, hippocampal, and cerebellar neurons, as well as in astrocytes (see image below).
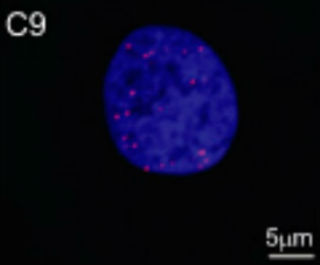
Spot the Antisense? A fluorescent probe for antisense transcripts labels RNA foci (red) in the nucleus of a fibroblast from a C9ORF72 expansion carrier. Image courtesy of National Academy of Sciences.
It is difficult to measure how much C9ORF72 expansion transcription goes on in the sense and antisense directions, researchers told Alzforum, because FISH probes for different transcripts may not be equally sensitive. Isaacs’ group found that cells from patients usually contained one or the other kind of RNA, though occasionally they spied cells containing both.
Do these foci differ in any physiological way? That is unknown. Tania Gendron of the Mayo Clinic in Jacksonville, Florida, pointed out that the degree of focus toxicity will likely depend on the particular RNA binding proteins it sequesters, which is yet to be determined. Gendron is a first author on another of the Acta Neuropathologica papers, together with senior authors Dennis Dickson and Leonard Petrucelli.
This group, and another led by Dieter Edbauer and Christian Haass at Ludwig-Maximilians University in Munich, wondered if the antisense RNAs could produce antisense peptides via a process called repeat-associated translation, which they had shown for C9ORF72 sense transcripts. To find out, the researchers engineered antibodies to the predicted antisense protein products. At Mayo, Gendron and joint first author Kevin Bieniek generated antibodies to the three peptides that could result from the three antisense reading frames: poly(proline-alanine), poly(proline-arginine), and poly(glycine-proline). Poly(GP) is also coded by the sense RNA expansion. Antibodies to poly(PA) and poly(PR) stained cytoplasmic inclusions in neurons of the hippocampus, motor cortex, temporal cortex, amygdala, thalamus, cerebellum, and medulla from C9ORF72 expansion carriers (see image below). They reported these results in the October 16 Acta Neuropathologica.
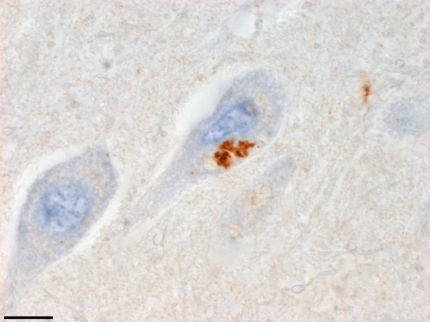
Antisense Makes Peptides. Antibodies to an antisense product of C9ORF72 expansion translation stain the hippocampus of a person who had FTD/ALS (scale bar=10 microns). Image courtesy of Tania Gendron and Kevin Bieniek, Mayo Clinic Jacksonville, Florida.
The researchers from Munich reported similar results in the same journal on October 17. Joint first authors Kohji Mori and Thomas Arzberger observed that antibodies to poly(PA) and poly(PR) stained cytoplasmic inclusions throughout the hippocampus and cortex, as well as in motor neurons. These two papers support a disease mechanism due to toxic peptides, the researchers agreed. Edbauer pointed out that either the peptide aggregates or the soluble peptide precursors could be toxic.
The five possible peptides appeared at different concentrations in the patient tissues. Edbauer and colleagues observed more of poly(GA) sense peptide, followed by poly(GR) sense peptide and poly(GP) sense/antisense. They saw less of the poly(PA) and poly(PR) antisense peptides. Edbauer suggested that the ratios could reflect different translation rates or stabilities of each peptide. Gendron said she and her colleagues are investigating the specific toxicity of each polypeptide, which she said may vary with such properties as the tendency to aggregate and to bind other proteins.
Sense and antisense C9ORF72 transcripts rarely exist in the same aggregate, but what about the five poly-dipeptides? The Edbauer group observed them mixed together in hippocampal cells taken from a person carrying the hexanucleotide expansion (see image below).
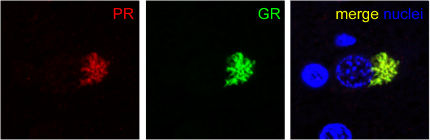
Sense Meets Antisense. Near a nucleus (blue), antibodies to the poly(PR) antisense peptide (red) and poly(GR) sense peptide (green) co-localize in the cytoplasm of cells from the hippocampus of a C9ORF72 repeat expansion carrier. Image courtesy of Dieter Edbauer and Kohji Mori, Ludwig-Maximilians University, Munich.
Thus far, no other repeat-expansion disorder has been shown to produce both sense and antisense RNA foci and peptides. The researchers who spoke with Alzforum knew of no other instance of expansions being translated into antisense peptide products. However, these new findings may inspire other scientists to look harder for bidirectional transcription and translation in other conditions, Ravits speculated.
Researchers next need to determine whether the expanded RNAs or the translated polypeptides are more toxic. Curiously, Gendron and colleagues noticed that cells from tissue samples of expansion carriers typically contain either RNA foci or peptide aggregates, but rarely both. Edbauer suggested that RNA incorporated into nuclear foci might be unavailable for translation.
Anti-sensibly Silencing C9ORF72
Whether toxic RNAs or toxic peptides are the problem, antisense oligonucleotide therapy might one day be a solution because such nucleotides tag transcripts for degradation by ribonuclease enzymes. Researchers already are testing this strategy clinically in ALS caused by mutated superoxide dismutase, and in Huntington’s disease, which is due to a trinucleotide expansion (see May 2013 news story and Jun 2012 news story). Earlier this month, two research groups reported that the strategy can detoxify RNA in cellular models of C9ORF72 disease (see news story and news story). Now, Ravits, co-senior author Don Cleveland, also at UCSD, and colleagues used antisense oligos to dissolve sense RNA foci in fibroblasts from people who carry C9ORF72 expansions. The treatment ablated expanded C9ORF72 isoforms while leaving alone the normal isoforms that make up the majority of C9ORF72 RNA, Ravits said. Mice treated with the antisense oligos appeared normal.
To understand what effects C9ORF72 might have on other genes, co-first authors Clotilde Lagier-Tourenne and Michael Baughn examined gene expression in fibroblasts from patients. They found 122 genes over-expressed and 34 under-expressed. Their oligonucleotide treatment dissolved the RNA foci but, surprisingly, the abnormal expression profile persisted. The authors posit that the treatment failed to correct gene expression because it left antisense RNA foci intact. Ravits suggested that dual therapy targeting both sense and antisense RNA may be necessary.
“I think the key finding from all these papers is that antisense RNA and repeat-associated translation might play a role in disease,” Isaacs said. That means scientists now have two RNAs and five peptides to investigate for potential binding partners and mechanisms of toxicity.
To cap off all this RNA research, the haploinsufficiency hypothesis also gained some currency this month. In the October 29 Acta Neuropathologica online, senior author Petrucelli and first authors Veronique Belzil and Peter Bauer, also at the Mayo Clinic, report that in the brains and blood of expansion carriers, the expanded C9ORF72 gene totes methylated histones. Such modifications are known to silence genes. Their results confirm a study that reported C9ORF72 methylation earlier this year (see Jun 2013 news story). The finding suggests that a partial loss of C9ORF72 function might also contribute to disease. Scientist have reported that in people with C9ORF72 repeat-based disease, expression of the C9ORF72 mRNA in blood and brain is abnormally low (see DeJesus-Hernandez et al., 2011, Gijselinck et al., 2012, and Renton et al., 2011).—Amber Dance
References
News Citations
- Corrupt Code: DNA Repeats Are Common Cause for ALS and FTD
- Chicago—RNA Inclusions Offer Therapeutic Target in ALS
- RNA Twist: C9ORF72 Intron Expansion Makes Aggregating Protein
- Second Study Sees Intron in FTLD Gene Translated
- Paper Alert: Antisense Oligonucleotide Therapy Safe for ALS?
- “Huntingtin Holiday” Helps Mice Back to Health
- RNA Deposits Confer Toxicity in C9ORF72 ALS
- Second Study Confirms Antisense Oligonucleotides Bust RNA Aggregates
- Methylation a Turn Off for Disease Gene C9ORF72?
Paper Citations
- Moseley ML, Zu T, Ikeda Y, Gao W, Mosemiller AK, Daughters RS, Chen G, Weatherspoon MR, Clark HB, Ebner TJ, Day JW, Ranum LP. Bidirectional expression of CUG and CAG expansion transcripts and intranuclear polyglutamine inclusions in spinocerebellar ataxia type 8. Nat Genet. 2006 Jul;38(7):758-69. PubMed.
- Ladd PD, Smith LE, Rabaia NA, Moore JM, Georges SA, Hansen RS, Hagerman RJ, Tassone F, Tapscott SJ, Filippova GN. An antisense transcript spanning the CGG repeat region of FMR1 is upregulated in premutation carriers but silenced in full mutation individuals. Hum Mol Genet. 2007 Dec 15;16(24):3174-87. PubMed.
- Chung DW, Rudnicki DD, Yu L, Margolis RL. A natural antisense transcript at the Huntington's disease repeat locus regulates HTT expression. Hum Mol Genet. 2011 Sep 1;20(17):3467-77. PubMed.
- Wilburn B, Rudnicki DD, Zhao J, Weitz TM, Cheng Y, Gu X, Greiner E, Park CS, Wang N, Sopher BL, La Spada AR, Osmand A, Margolis RL, Sun YE, Yang XW. An Antisense CAG Repeat Transcript at JPH3 Locus Mediates Expanded Polyglutamine Protein Toxicity in Huntington's Disease-like 2 Mice. Neuron. 2011 May 12;70(3):427-40. PubMed.
- DeJesus-Hernandez M, Mackenzie IR, Boeve BF, Boxer AL, Baker M, Rutherford NJ, Nicholson AM, Finch NA, Flynn H, Adamson J, Kouri N, Wojtas A, Sengdy P, Hsiung GY, Karydas A, Seeley WW, Josephs KA, Coppola G, Geschwind DH, Wszolek ZK, Feldman H, Knopman DS, Petersen RC, Miller BL, Dickson DW, Boylan KB, Graff-Radford NR, Rademakers R. Expanded GGGGCC hexanucleotide repeat in noncoding region of C9ORF72 causes chromosome 9p-linked FTD and ALS. Neuron. 2011 Oct 20;72(2):245-56. Epub 2011 Sep 21 PubMed.
- Gijselinck I, Van Langenhove T, van der Zee J, Sleegers K, Philtjens S, Kleinberger G, Janssens J, Bettens K, Van Cauwenberghe C, Pereson S, Engelborghs S, Sieben A, De Jonghe P, Vandenberghe R, Santens P, De Bleecker J, Maes G, Bäumer V, Dillen L, Joris G, Cuijt I, Corsmit E, Elinck E, Van Dongen J, Vermeulen S, Van den Broeck M, Vaerenberg C, Mattheijssens M, Peeters K, Robberecht W, Cras P, Martin JJ, De Deyn PP, Cruts M, Van Broeckhoven C. A C9orf72 promoter repeat expansion in a Flanders-Belgian cohort with disorders of the frontotemporal lobar degeneration-amyotrophic lateral sclerosis spectrum: a gene identification study. Lancet Neurol. 2012 Jan;11(1):54-65. PubMed.
- Renton AE, Majounie E, Waite A, Simón-Sánchez J, Rollinson S, Gibbs JR, Schymick JC, Laaksovirta H, van Swieten JC, Myllykangas L, Kalimo H, Paetau A, Abramzon Y, Remes AM, Kaganovich A, Scholz SW, Duckworth J, Ding J, Harmer DW, Hernandez DG, Johnson JO, Mok K, Ryten M, Trabzuni D, Guerreiro RJ, Orrell RW, Neal J, Murray A, Pearson J, Jansen IE, Sondervan D, Seelaar H, Blake D, Young K, Halliwell N, Callister JB, Toulson G, Richardson A, Gerhard A, Snowden J, Mann D, Neary D, Nalls MA, Peuralinna T, Jansson L, Isoviita VM, Kaivorinne AL, Hölttä-Vuori M, Ikonen E, Sulkava R, Benatar M, Wuu J, Chiò A, Restagno G, Borghero G, Sabatelli M, ITALSGEN Consortium, Heckerman D, Rogaeva E, Zinman L, Rothstein JD, Sendtner M, Drepper C, Eichler EE, Alkan C, Abdullaev Z, Pack SD, Dutra A, Pak E, Hardy J, Singleton A, Williams NM, Heutink P, Pickering-Brown S, Morris HR, Tienari PJ, Traynor BJ. A hexanucleotide repeat expansion in C9ORF72 is the cause of chromosome 9p21-linked ALS-FTD. Neuron. 2011 Oct 20;72(2):257-68. Epub 2011 Sep 21 PubMed.
Further Reading
Papers
- Todd PK, Paulson HL. C9orf72-Associated FTD/ALS: When Less Is More. Neuron. 2013 Oct 16;80(2):257-8. PubMed.
- Cleary JD, Ranum LP. Repeat-associated non-ATG (RAN) translation in neurological disease. Hum Mol Genet. 2013 Oct 15;22(R1):R45-51. PubMed.
- Dols-Icardo O, García-Redondo A, Rojas-García R, Sánchez-Valle R, Noguera A, Gómez-Tortosa E, Pastor P, Hernández I, Esteban-Pérez J, Suárez-Calvet M, Antón-Aguirre S, Amer G, Ortega-Cubero S, Blesa R, Fortea J, Alcolea D, Capdevila A, Antonell A, Lladó A, Muñoz-Blanco JL, Mora JS, Galán-Dávila L, Rodríguez de Rivera FJ, Lleó A, Clarimón J. Characterization of the repeat expansion size in C9orf72 in amyotrophic lateral sclerosis and frontotemporal dementia. Hum Mol Genet. 2013 Oct 8; PubMed.
- Gendron TF, Cosio DM, Petrucelli L. c9RAN translation: a potential therapeutic target for the treatment of amyotrophic lateral sclerosis and frontotemporal dementia. Expert Opin Ther Targets. 2013 Sep;17(9):991-5. PubMed.
- Belzil VV, Gendron TF, Petrucelli L. RNA-mediated toxicity in neurodegenerative disease. Mol Cell Neurosci. 2012 Dec 29; PubMed.
- Mori K, Lammich S, Mackenzie IR, Forné I, Zilow S, Kretzschmar H, Edbauer D, Janssens J, Kleinberger G, Cruts M, Herms J, Neumann M, Van Broeckhoven C, Arzberger T, Haass C. hnRNP A3 binds to GGGGCC repeats and is a constituent of p62-positive/TDP43-negative inclusions in the hippocampus of patients with C9orf72 mutations. Acta Neuropathol. 2013 Mar;125(3):413-23. PubMed.
News
- Brain Imaging Distinguishes C9ORF72 From Other Types of ALS
- Paper Alert: Antisense Oligonucleotide Therapy Safe for ALS?
- C9ORF72 Function: Is the ALS Protein a Membrane Traffic Cop?
- Chicago—Dynamic Repeats: C9ORF72 Expands and Shrinks in ALS
- C9ORF72 Steals the Show at Frontotemporal Dementia Meeting
- When Is a C9ORF72 Repeat Expansion Not a C9ORF72 Repeat Expansion?
Primary Papers
- Lagier-Tourenne C, Baughn M, Rigo F, Sun S, Liu P, Li HR, Jiang J, Watt AT, Chun S, Katz M, Qiu J, Sun Y, Ling SC, Zhu Q, Polymenidou M, Drenner K, Artates JW, McAlonis-Downes M, Markmiller S, Hutt KR, Pizzo DP, Cady J, Harms MB, Baloh RH, Vandenberg SR, Yeo GW, Fu XD, Bennett CF, Cleveland DW, Ravits J. Targeted degradation of sense and antisense C9orf72 RNA foci as therapy for ALS and frontotemporal degeneration. Proc Natl Acad Sci U S A. 2013 Nov 19;110(47):E4530-9. Epub 2013 Oct 29 PubMed.
- Belzil VV, Bauer PO, Prudencio M, Gendron TF, Stetler CT, Yan IK, Pregent L, Daughrity L, Baker MC, Rademakers R, Boylan K, Patel TC, Dickson DW, Petrucelli L. Reduced C9orf72 gene expression in c9FTD/ALS is caused by histone trimethylation, an epigenetic event detectable in blood. Acta Neuropathol. 2013 Dec;126(6):895-905. Epub 2013 Oct 29 PubMed.
- Mizielinska S, Lashley T, Norona FE, Clayton EL, Ridler CE, Fratta P, Isaacs AM. C9orf72 frontotemporal lobar degeneration is characterised by frequent neuronal sense and antisense RNA foci. Acta Neuropathol. 2013 Oct 30; PubMed.
- Gendron TF, Bieniek KF, Zhang YJ, Jansen-West K, Ash PE, Caulfield T, Daughrity L, Dunmore JH, Castanedes-Casey M, Chew J, Cosio DM, van Blitterswijk M, Lee WC, Rademakers R, Boylan KB, Dickson DW, Petrucelli L. Antisense transcripts of the expanded C9ORF72 hexanucleotide repeat form nuclear RNA foci and undergo repeat-associated non-ATG translation in c9FTD/ALS. Acta Neuropathol. 2013 Oct 16; PubMed.
- Mori K, Arzberger T, Grässer FA, Gijselinck I, May S, Rentzsch K, Weng SM, Schludi MH, van der Zee J, Cruts M, Van Broeckhoven C, Kremmer E, Kretzschmar HA, Haass C, Edbauer D. Bidirectional transcripts of the expanded C9orf72 hexanucleotide repeat are translated into aggregating dipeptide repeat proteins. Acta Neuropathol. 2013 Oct 17; PubMed. Correction.
Annotate
To make an annotation you must Login or Register.

Comments
No Available Comments
Make a Comment
To make a comment you must login or register.