Mutations
MAPT IVS10+16 C>T
Quick Links
Overview
Pathogenicity: Frontotemporal Dementia : Pathogenic
Clinical
Phenotype: Alzheimer's Disease, Frontotemporal Dementia, Progressive Supranuclear Palsy
Position: (GRCh38/hg38):Chr17:46010418 C>T
Position: (GRCh37/hg19):Chr17:44087784 C>T
dbSNP ID: rs63751011
Coding/Non-Coding: Non-Coding
DNA
Change: Substitution
Expected RNA
Consequence: Splicing Alteration
Expected Protein
Consequence: Isoform Shift
Codon
Change: C to T
Genomic
Region: Intron 10
Research
Models: 2
Findings
This is a relatively common MAPT mutation found in families from around the world. Individuals with this mutation display a variety of phenotypes and have been diagnosed with frontotemporal dementia, Alzheimer's disease, progressive supranuclear palsy, progressive subcortical gliosis, and frontotemporal dementia with parkinsonism.
This intronic mutation was originally found in four families (one from Australia, one from the United States, and two from the United Kingdom), two of which had prior evidence of genetic linkage to chromosome 17. The Australian kindred, known as Aus1, had 28 affected members over five generations with a mean age at onset of 53. A U.S. kindred, known as FTD002, had three affected members over two generations with a mean age at onset of 40. The U.K. kindreds, known as Man6 and Man23, had two and 10 affected members in one and two generations, with mean onsets at 48 and 51 years old, respectively. In each family the mutation segregated with disease (Hutton et al., 1998).
Neuropathology
Four autopsied cases in the Australian kindred showed neuronal loss, gray-matter gliosis, and neuropil vacuolation in both frontal and temporal lobes. Balloon neurons in the cortex were also observed, along with degeneration of the substantia nigra with free melanin. Tau-positive neurons were observed in select brain regions. Amyloid plaques, Lewy bodies, and Pick bodies were not observed (Baker et al., 1997).
Biological Effect
This intronic mutation destabilizes a stem-loop structure that regulates the alternative splicing of exon 10. This causes more frequent inclusion of exon 10 and an increased proportion of four-repeat (4R) tau isoforms (Hutton et al., 1998).
Induced pluripotent stem cells (iPSCs) have been generated from the fibroblasts of two FTD patients with this mutation (Sposito et al., 2015). Cortical neurons derived from the iPSCs express both 0N3R and 0N4R isoforms of tau. In contrast, equivalent cells derived from control individuals display a splicing pattern more consistent with known developmental regulation of tau (i.e. 0N3R developmentally followed by a mix of 3R and 4R isoforms), suggesting that the 10+16 mutation may override the normal developmental pattern of tau splicing.
Research Models
A transgenic mouse model carrying a human transgene with this intronic mutation recapitulates many disease-related phenotypes, including abnormal tau phosphorylation, synapse loss and dysfunction, memory impairment, glial activation, tangle formation, and age-dependent neuronal loss. The model, called Tau609, also has an imbalance of 3R and 4R tau isoforms (higher 4R), confirming in vitro work implicating this intronic mutation in the misregulation of exon 10 splicing (Umeda et al., 2013).
Last Updated: 18 Jul 2024
References
Research Models Citations
Paper Citations
- Umeda T, Yamashita T, Kimura T, Ohnishi K, Takuma H, Ozeki T, Takashima A, Tomiyama T, Mori H. Neurodegenerative disorder FTDP-17-related tau intron 10 +16C → T mutation increases tau exon 10 splicing and causes tauopathy in transgenic mice. Am J Pathol. 2013 Jul;183(1):211-25. PubMed.
- Hutton M, Lendon CL, Rizzu P, Baker M, Froelich S, Houlden H, Pickering-Brown S, Chakraverty S, Isaacs A, Grover A, Hackett J, Adamson J, Lincoln S, Dickson D, Davies P, Petersen RC, Stevens M, de Graaff E, Wauters E, van Baren J, Hillebrand M, Joosse M, Kwon JM, Nowotny P, Che LK, Norton J, Morris JC, Reed LA, Trojanowski J, Basun H, Lannfelt L, Neystat M, Fahn S, Dark F, Tannenberg T, Dodd PR, Hayward N, Kwok JB, Schofield PR, Andreadis A, Snowden J, Craufurd D, Neary D, Owen F, Oostra BA, Hardy J, Goate A, van Swieten J, Mann D, Lynch T, Heutink P. Association of missense and 5'-splice-site mutations in tau with the inherited dementia FTDP-17. Nature. 1998 Jun 18;393(6686):702-5. PubMed.
- Baker M, Kwok JB, Kucera S, Crook R, Farrer M, Houlden H, Isaacs A, Lincoln S, Onstead L, Hardy J, Wittenberg L, Dodd P, Webb S, Hayward N, Tannenberg T, Andreadis A, Hallupp M, Schofield P, Dark F, Hutton M. Localization of frontotemporal dementia with parkinsonism in an Australian kindred to chromosome 17q21-22. Ann Neurol. 1997 Nov;42(5):794-8. PubMed.
- Sposito T, Preza E, Mahoney CJ, Setó-Salvia N, Ryan NS, Morris HR, Arber C, Devine MJ, Houlden H, Warner TT, Bushell TJ, Zagnoni M, Kunath T, Livesey FJ, Fox NC, Rossor MN, Hardy J, Wray S. Developmental regulation of tau splicing is disrupted in stem cell-derived neurons from frontotemporal dementia patients with the 10 + 16 splice-site mutation in MAPT. Hum Mol Genet. 2015 Sep 15;24(18):5260-9. Epub 2015 Jul 1 PubMed.
Further Reading
Papers
- Houlden H, Baker M, Adamson J, Grover A, Waring S, Dickson D, Lynch T, Boeve B, Petersen RC, Pickering-Brown S, Owen F, Neary D, Craufurd D, Snowden J, Mann D, Hutton M. Frequency of tau mutations in three series of non-Alzheimer's degenerative dementia. Ann Neurol. 1999 Aug;46(2):243-8. PubMed.
- Poorkaj P, Grossman M, Steinbart E, Payami H, Sadovnick A, Nochlin D, Tabira T, Trojanowski JQ, Borson S, Galasko D, Reich S, Quinn B, Schellenberg G, Bird TD. Frequency of tau gene mutations in familial and sporadic cases of non-Alzheimer dementia. Arch Neurol. 2001 Mar;58(3):383-7. PubMed.
- Morris HR, Khan MN, Janssen JC, Brown JM, Perez-Tur J, Baker M, Ozansoy M, Hardy J, Hutton M, Wood NW, Lees AJ, Revesz T, Lantos P, Rossor MN. The genetic and pathological classification of familial frontotemporal dementia. Arch Neurol. 2001 Nov;58(11):1813-6. PubMed.
- Pickering-Brown SM, Richardson AM, Snowden JS, McDonagh AM, Burns A, Braude W, Baker M, Liu WK, Yen SH, Hardy J, Hutton M, Davies Y, Allsop D, Craufurd D, Neary D, Mann DM. Inherited frontotemporal dementia in nine British families associated with intronic mutations in the tau gene. Brain. 2002 Apr;125(Pt 4):732-51. PubMed.
- Rohrer JD, Guerreiro R, Vandrovcova J, Uphill J, Reiman D, Beck J, Isaacs AM, Authier A, Ferrari R, Fox NC, Mackenzie IR, Warren JD, de Silva R, Holton J, Revesz T, Hardy J, Mead S, Rossor MN. The heritability and genetics of frontotemporal lobar degeneration. Neurology. 2009 Nov 3;73(18):1451-6. PubMed.
- Rohrer JD, Ridgway GR, Modat M, Ourselin S, Mead S, Fox NC, Rossor MN, Warren JD. Distinct profiles of brain atrophy in frontotemporal lobar degeneration caused by progranulin and tau mutations. Neuroimage. 2010 Nov 15;53(3):1070-6. Epub 2010 Jan 4 PubMed.
- Stanford PM, Brooks WS, Teber ET, Hallupp M, McLean C, Halliday GM, Martins RN, Kwok JB, Schofield PR. Frequency of tau mutations in familial and sporadic frontotemporal dementia and other tauopathies. J Neurol. 2004 Sep;251(9):1098-104. PubMed.
- Janssen JC, Warrington EK, Morris HR, Lantos P, Brown J, Revesz T, Wood N, Khan MN, Cipolotti L, Fox NC, Rossor MN. Clinical features of frontotemporal dementia due to the intronic tau 10(+16) mutation. Neurology. 2002 Apr 23;58(8):1161-8. PubMed.
- Tsuboi Y, Uitti RJ, Baker M, Hutton ML, Wszolek ZK. Clinical features of frontotemporal dementia due to the intronic tau 10(+16) mutation. Neurology. 2003 Feb 11;60(3):525-6. PubMed.
- Takada LT, Bahia VS, Guimarães HC, Costa TV, Vale TC, Rodriguez RD, Porto FH, Machado JC, Beato RG, Cesar KG, Smid J, Nascimento CF, Grinberg LT, Brucki SM, Maximino JR, Camargos ST, Chadi G, Caramelli P, Nitrini R. GRN and MAPT Mutations in 2 Frontotemporal Dementia Research Centers in Brazil. Alzheimer Dis Assoc Disord. 2016 Apr 14; PubMed.
Learn More
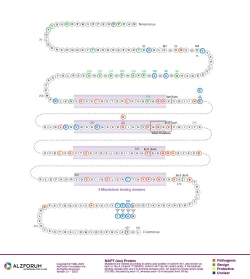
Protein Diagram
Primary Papers
- Hutton M, Lendon CL, Rizzu P, Baker M, Froelich S, Houlden H, Pickering-Brown S, Chakraverty S, Isaacs A, Grover A, Hackett J, Adamson J, Lincoln S, Dickson D, Davies P, Petersen RC, Stevens M, de Graaff E, Wauters E, van Baren J, Hillebrand M, Joosse M, Kwon JM, Nowotny P, Che LK, Norton J, Morris JC, Reed LA, Trojanowski J, Basun H, Lannfelt L, Neystat M, Fahn S, Dark F, Tannenberg T, Dodd PR, Hayward N, Kwok JB, Schofield PR, Andreadis A, Snowden J, Craufurd D, Neary D, Owen F, Oostra BA, Hardy J, Goate A, van Swieten J, Mann D, Lynch T, Heutink P. Association of missense and 5'-splice-site mutations in tau with the inherited dementia FTDP-17. Nature. 1998 Jun 18;393(6686):702-5. PubMed.
Alzpedia
Disclaimer: Alzforum does not provide medical advice. The Content is for informational, educational, research and reference purposes only and is not intended to substitute for professional medical advice, diagnosis or treatment. Always seek advice from a qualified physician or health care professional about any medical concern, and do not disregard professional medical advice because of anything you may read on Alzforum.

Comments
No Available Comments
Make a Comment
To make a comment you must login or register.