Massive Proteomics Study Connects Genes, Proteins, Disease
Quick Links
Genome-wide association studies have yielded astronomical amounts of data connecting genetic variation to disease. Yet the millions of SNPs identified in GWAS do not by themselves explain how disease develops. Proteins—the end product of gene expression—lie closer to the mechanistic meat. To illuminate the links between SNPs, proteins, and disease, researchers led by Kari Stefansson of deCODE Genetics in Reykjavik, surveyed more proteins, in more people, than anyone has even done before. As described December 2 in Nature Genetics, they measured levels of nearly 5,000 proteins in the plasma of more than 35,000 people.
- Largest plasma proteomics study to date probed 5,000 proteins in 35,000 people.
- Identifies 18,000+ protein quantitative trait loci, or pQTLs.
- Connects dots between protein levels, genetic variation, and disease.
- Variants at MS4A locus dial down soluble TREM2 in plasma.
Integrating genomic data on the same participants, they detected more than 18,000 protein quantitative trait loci, or pQTLs—genetic variants that correlate with the plasma level of protein. Weaving in data from GWAS, they then connected the dots from genetic variation to protein to disease, confirming known causal connections or uncovering new ones. One was none other than TREM2. The authors found that reduced plasma levels of the soluble extracellular domain of the microglial receptor correlated with AD risk variants near the MS4A gene, confirming previous reports that variants at this locus sway AD risk via TREM2.
“This study is a massive undertaking in relating genetic variants to protein levels in plasma,” commented Betty Tijms of Amsterdam University Medical Center and Pieter Jelle Visser of Maastricht University in the Netherlands. “Proteomics measured in biofluids provides impelling opportunities to study in great detail how genetic risk factors impact on disease pathogenesis in patients.”
“This data will be instrumental in identifying functional and causal genes and proteins for complex traits and diseases such as AD,” said Carlos Cruchaga of Washington University in St. Louis. “We’re going to learn a lot of new biology.”
The study is not the first of its kind, but it is, by far, the largest. One previous pQTL study included almost as many participants but surveyed only 90 proteins, while the largest study to measure at least 200 proteins included fewer than 6,000 people (Emilsson et al., 2018; Folkersen et al., 2020). Another recent study analyzed more than 8,000 proteins in brain samples from 376 people (Feb 2021news).
First author Egil Ferkingstad and colleagues used aptamers targeting 4,907 proteins to survey the plasma proteomes of 35,559 Icelanders, each of whom also had had either whole-genome sequencing or genotyping done as part of the Icelandic Cancer Project or of various other genetics projects at deCODE. Among 27.2 million genetic sequence variants detected within this population, the scientists spotted 18,084 that correlated with plasma levels of a protein. Of these, 1,881 were cis-pQTLs, meaning that the sequence variant resided in or near the gene encoding the associated protein. The remaining 16,203 were trans-pQTLs, landing in a disparate region of the genome from the gene encoding the associated protein. This trove upped the number of pQTLs discovered by all previous studies by an order of magnitude.
Next, the researchers brought a wealth of phenotypic information—collected for all participants—into the fold. They tested for associations between plasma protein levels and a set of 373 diseases and other traits, ultimately digging up a whopping 257,490 links between protein levels and phenotypes.
An association between the level of a circulating protein and a disease cannot distinguish whether the altered protein level is a cause or consequence of the disease process. To get closer to filling in that blank, Cruchaga said, one needs a third piece of information: associated genetic variation. To get this, the researchers integrated GWAS data into their analysis. Scanning the NHGRI-EBI catalog of GWAS hits, the researchers identified 45,334 variants that associated with diseases and traits. Of these, 5,458—or 12 percent—overlapped, or were co-inherited, with at least one pQTL identified in the Icelandic cohort. This suggested that these disease risk variants also correlated with plasma protein levels associated with their respective pQTL.
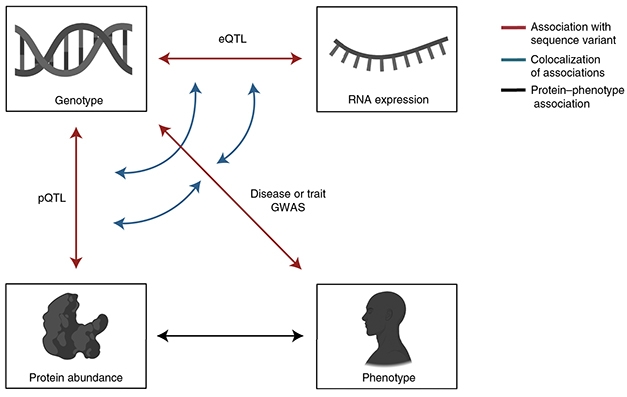
Integrating 'Omics. Associations between genetic sequence variants and protein level, RNA expression, and phenotypes are shown as red arrows. Co-localizations of associations are shown as blue arrows. [Courtesy of Ferkingstad et al., Nature Genetics, 2021.]
What were these troikas of GWAS, pQTLs, and plasma protein level? One example involved the AD risk variant, TREM2. The R47H variant within the TREM2 gene is known to raise AD risk, but a previous study demonstrated that a variant close to the MS4A gene cluster not only influenced AD risk, but also associated with reduced levels of soluble TREM2 in the cerebrospinal fluid (Jul 2018 news). Lo and behold, Ferkingstad confirmed this in the pQTL study, identifying the same AD risk variant as a trans-pQTL that associated with reduced plasma sTREM2. The findings strengthen the idea that MS4A variants influence AD risk by modulating sTREM2 levels.

TREM2 Connection. Around the MS4A gene cluster (bottom) many variants associate with AD (middle). Variants at the same locus also associate with reduced plasma sTREM2 (top). [Courtesy of Ferkingstad et al., Nature Genetics, 2021.]
While TREM2 is a select highlight for Alzheimerologists, the scientists scoured their heap of data to clarify other disease associations. For example, a variant that reduces a person's risk for colorectal cancer and lies near the CHRDL2 gene associated with lower plasma levels of the CHRDL2 protein, which antagonizes bone morphogenetic protein (BMP), a member of the transforming growth factor β superfamily. Likewise, another colorectal cancer risk reducing variant near the gene encoding a different BMP antagonist, gremlin-1, also associated with lower levels of gremlin-1 in plasma. Together, these findings not only nail the causal genes associated with the risk variants, but also provide support for the idea that BMP signaling protects against this type of cancer.
The findings may help researchers discover plasma biomarkers. Case in point: one rare genetic variant in the transthyretin (TTR) gene causes a hereditary form of amyloidosis that manifests with polyneuropathy, while another variant in the same gene protects against the disease. In their dataset, the scientists found that people with the disease variant had higher plasma concentrations of ADP ribosylation factor 3, while people with the protective variant had reduced levels of TNF receptor superfamily 3, encoded by the NGFR gene. They propose that these proteins—with opposite relationships to disease—are potential biomarkers of treatment response and progression of hereditary TTR amyloidosis.
Overall, the findings underscore that protein levels are highly regulated by genetic variants, Cruchaga said. While some of these proteins—such as soluble TREM2—wind up in the blood, others may remain sequestered in the brain. Herein lies a limitation of this plasma proteomics study, he noted, as it will not pinpoint genetic variants that regulate proteins that never make it to the plasma.—Jessica Shugart
References
News Citations
- PWAS x GWAS? Proteome Analysis Nets 10 New Alzheimer’s Genes
- MS4A Alzheimer’s Risk Gene Linked to TREM2 Signaling
Paper Citations
- Emilsson V, Ilkov M, Lamb JR, Finkel N, Gudmundsson EF, Pitts R, Hoover H, Gudmundsdottir V, Horman SR, Aspelund T, Shu L, Trifonov V, Sigurdsson S, Manolescu A, Zhu J, Olafsson Ö, Jakobsdottir J, Lesley SA, To J, Zhang J, Harris TB, Launer LJ, Zhang B, Eiriksdottir G, Yang X, Orth AP, Jennings LL, Gudnason V. Co-regulatory networks of human serum proteins link genetics to disease. Science. 2018 Aug 24;361(6404):769-773. Epub 2018 Aug 2 PubMed.
- Folkersen L, Gustafsson S, Wang Q, Hansen DH, Hedman ÅK, Schork A, Page K, Zhernakova DV, Wu Y, Peters J, Eriksson N, Bergen SE, Boutin TS, Bretherick AD, Enroth S, Kalnapenkis A, Gådin JR, Suur BE, Chen Y, Matic L, Gale JD, Lee J, Zhang W, Quazi A, Ala-Korpela M, Choi SH, Claringbould A, Danesh J, Davey Smith G, de Masi F, Elmståhl S, Engström G, Fauman E, Fernandez C, Franke L, Franks PW, Giedraitis V, Haley C, Hamsten A, Ingason A, Johansson Å, Joshi PK, Lind L, Lindgren CM, Lubitz S, Palmer T, Macdonald-Dunlop E, Magnusson M, Melander O, Michaelsson K, Morris AP, Mägi R, Nagle MW, Nilsson PM, Nilsson J, Orho-Melander M, Polasek O, Prins B, Pålsson E, Qi T, Sjögren M, Sundström J, Surendran P, Võsa U, Werge T, Wernersson R, Westra HJ, Yang J, Zhernakova A, Ärnlöv J, Fu J, Smith JG, Esko T, Hayward C, Gyllensten U, Landen M, Siegbahn A, Wilson JF, Wallentin L, Butterworth AS, Holmes MV, Ingelsson E, Mälarstig A. Genomic and drug target evaluation of 90 cardiovascular proteins in 30,931 individuals. Nat Metab. 2020 Oct;2(10):1135-1148. Epub 2020 Oct 16 PubMed.
External Citations
Further Reading
No Available Further Reading
Primary Papers
- Ferkingstad E, Sulem P, Atlason BA, Sveinbjornsson G, Magnusson MI, Styrmisdottir EL, Gunnarsdottir K, Helgason A, Oddsson A, Halldorsson BV, Jensson BO, Zink F, Halldorsson GH, Masson G, Arnadottir GA, Katrinardottir H, Juliusson K, Magnusson MK, Magnusson OT, Fridriksdottir R, Saevarsdottir S, Gudjonsson SA, Stacey SN, Rognvaldsson S, Eiriksdottir T, Olafsdottir TA, Steinthorsdottir V, Tragante V, Ulfarsson MO, Stefansson H, Jonsdottir I, Holm H, Rafnar T, Melsted P, Saemundsdottir J, Norddahl GL, Lund SH, Gudbjartsson DF, Thorsteinsdottir U, Stefansson K. Large-scale integration of the plasma proteome with genetics and disease. Nature Genetics, December 2, 2021
Annotate
To make an annotation you must Login or Register.

Comments
Amsterdam UMC, loc. VUmc
Maastricht University; VU University Medical Centre
This study is a massive undertaking, relating genetic variants to protein levels in plasma. Proteomics measured in biofluids provides impelling opportunities to study in great detail how genetic risk factors impact on disease pathogenesis in patients.
Ferkingstad et al. used this approach and measured in existing blood samples proteomics with the aptamer approach of Somascan in 35,559 individuals, which is almost 15 percent of the total population of Iceland. Note that part of these samples were enriched for cancer as they were acquired in the context of the Icelandic Cancer project.
About a quarter of proteins showed cis associations, meaning that a variant in a given gene was related to plasma concentrations of the corresponding protein. The majority of associations were, however, trans, with a given variant influencing concentrations of another protein than it encodes. One of those trans associations was a variant cluster surrounding the MS4A6A and MSA4A genes, which are expressed in microglia. Variants in those genes have been associated with Alzheimer’s disease before, and in this study were associated with lower plasma concentrations of TREM2, replicating another study (Deming et al., 2019).
TREM2 itself also has known AD risk variants. The authors do not go into detail on those relationships, although they suggest that certain variants may lead to artefactual measures of plasma TREM2 with Somascan. A previous study comparing rare TREM2 variants on plasma TREM2 levels did not find clear associations (Ashton et al., 2019). Together these results further support involvement of microglia in AD disease pathogenesis. However, from the main text it was unclear whether the presence of AD pathology in individuals carrying the variants was assessed, as well as their clinical diagnosis and age.
In the case of Alzheimer’s disease, it would be of interest to measure plasma p-tau levels as well to further understand which gene-protein concentrations may be related to amyloid pathology in the brain. Finally, it would be interesting to study whether gene-protein associations would be similar for cerebrospinal fluid.
References:
Deming Y, Filipello F, Cignarella F, Cantoni C, Hsu S, Mikesell R, Li Z, Del-Aguila JL, Dube U, Farias FG, Bradley J, Budde J, Ibanez L, Fernandez MV, Blennow K, Zetterberg H, Heslegrave A, Johansson PM, Svensson J, Nellgård B, Lleo A, Alcolea D, Clarimon J, Rami L, Molinuevo JL, Suárez-Calvet M, Morenas-Rodríguez E, Kleinberger G, Ewers M, Harari O, Haass C, Brett TJ, Benitez BA, Karch CM, Piccio L, Cruchaga C. The MS4A gene cluster is a key modulator of soluble TREM2 and Alzheimer's disease risk. Sci Transl Med. 2019 Aug 14;11(505) PubMed.
Ashton NJ, Suárez-Calvet M, Heslegrave A, Hye A, Razquin C, Pastor P, Sanchez-Valle R, Molinuevo JL, Visser PJ, Blennow K, Hodges AK, Zetterberg H. Plasma levels of soluble TREM2 and neurofilament light chain in TREM2 rare variant carriers. Alzheimers Res Ther. 2019 Nov 28;11(1):94. PubMed.
Make a Comment
To make a comment you must login or register.