5,000 Down, 95,000 to Go: U.K. Biobank Releases First Brain-Imaging Data
Quick Links
Inside a nondescript building just south of Manchester, England, a new person squeezes into an MRI tube every 36 minutes. This machine subserves a single purpose: to conduct the largest imaging study in human history. The brain-scanning portion of the U.K. Biobank epidemiological study aims to image 100,000 healthy people within five years. The goal is to uncover early warning signs of neurological and other diseases. In the September 19 Nature Neuroscience, the researchers, led by Stephen Smith at the University of Oxford in England, published data from the first 5,000 volunteers scanned.
This initial release appeared to be a practice run at the formidable task of distilling massive amounts of complex imaging data into digestible findings. The researchers made connections between the structure and function of different brain regions on the one hand, and a myriad of health and lifestyle measures for each participant that are also being recorded on the other. While this initial analysis only looked for baseline correlations, the real meat of the project will come later, when people in the cohort start to develop diseases such as AD. Then the researchers will re-examine the imaging and health data to dig for early harbingers of disease. Ultimately, other researchers who tap this open-access data in the future can analyze it however they choose, but the present paper is a first stab at parsing this ever-growing data set.
“The ambition to generate the quantity of imaging data described in this paper is simply phenomenal, and this description of the process and results from the first 5,000 scans marks a step change in imaging sciences,” commented Simon Lovestone, also from the University of Oxford. While not involved with the current paper, Lovestone plans to utilize U.K. Biobank data. “Given that the U.K. Biobank data, including the imaging data reported here, is readily available within the spirit of open access, its uses will be limited only by the scientific imagination of the research community,” he wrote. Other AD scientists note that it is also limited by not testing for known molecular pathology markers, which could prove invaluable later on for determining when AD started, in whom, and why.
U.K. Biobank is a broad prospective epidemiological study that aims to uncover connections between various diseases and health and lifestyle factors that precede them. Researchers are collecting extensive phenotypic data, using exhaustive health and lifestyle questionnaires, whole body and cardiac imaging, blood samples, genetic testing, and medical records.
Its scope is massive. Half a million participants, aged 40-69 years, will be tracked for the emergence of a variety of health outcomes, including neurodegenerative disease. Of these volunteers, 100,000 will also undergo structural, functional, and diffusion MRI of the brain (described in the current study), whole body and heart imaging also by MRI, low-dose X-rays of bone and joints, and ultrasound of the carotid arteries. The researchers packed as many different brain imaging modalities as possible into a 36-minute window to allow them to scan 18 participants per day, seven days a week, and complete the imaging phase by 2022.
Notably, participants will not undergo PET scans of any kind, an omission some neurodegenerative disease researchers lament. Because PET scans can capture Aβ and tau pathology, which precede structural changes in the brain and the appearance of cognitive symptoms, their usefulness in studying presymptomatic Alzheimer’s and other tauopathies is matched only by CSF analysis. The U.K. Biobank will not collect CSF either.
While this first release covers 5,000 participants, first author Karla Miller told Alzforum that as of this writing, more than 11,000 people have been scanned in Manchester, with two more locations coming online soon. Costs for the brain-imaging portion of the study—funded by the U.K. Medical Research Council and Wellcome Trust—are projected to reach £40 million, Miller said.
What does the first batch of data show? Taken together, different types of structural MRI revealed the basic shape and size of different parts of the brain, the appearance of white-matter lesions, and deposits of iron, which are associated with aging and neurodegeneration. Diffusion MRI revealed the integrity of microstructures in the brain, including tracts of white matter that connect different regions. Finally, neural activity and connections between different brain regions were assessed using functional MRI, conducted as participants rested or were immersed in a particular task.
In an attempt to make biological sense out of this sprawling mass of data, Miller and colleagues drew all the imaging data down to 2,501 image-derived phenotypes (IDPs). Each IDP represents a single parameter, such as the volume of a specific brain region, the connection strength between two parts of the brain, or the level of neural activity that flares up when a participant completes a certain task. The researchers then looked for associations between these IDPs and other health data stored in the Biobank. The researchers divided these 1,100 health data measures into 11 categories, which included lifestyle factors such as exercise and diet, physical attributes such as body-mass index and bone density, and scores on cognitive tests.
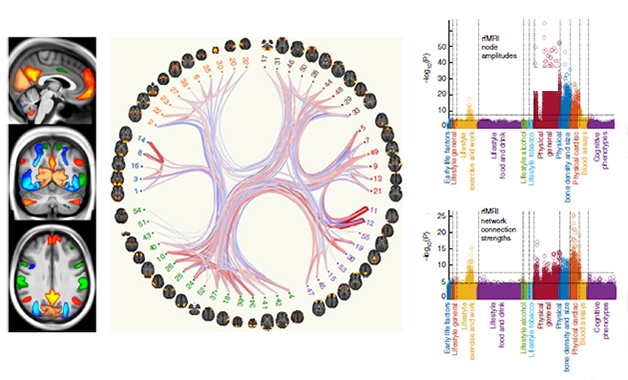
Divvying Up Deep Data. The strength of thousands of connections (middle) between different brain networks (left) could be correlated to various health-related phenotypes (right). [Image courtesy of Miller et al., Nature Neuroscience, 2016.]
After adjusting for age, sex, and head size, the researchers found strong correlations between various IDPs and other measures. For example, performance on the digit-symbol substitution test, which wanes in people with dementia, correlated with thalamic volume as well as the integrity of three white-matter tracts: the corona radiata, the superior thalamic radiation, and the posterior fornix. Scores on a reaction-time test correlated with the size of the left putamen, while fluid intelligence scores correlated with the strength of neural activity when participants performed a simple shape-matching task. Neurodegeneration in the amygdala and hippocampus—as inferred from elevated iron deposition in those structures—correlated with elevated body-mass index. This latter association confirmed what previously had been reported in the Austrian Stroke Prevention Study, suggesting the U.K. Biobank data set can corroborate previous findings (see Pirpamer et al., 2016).
What do correlations like these really mean? Miller said it’s impossible to know at this point, and that researchers should use the data to form hypotheses for testing, rather than drawing causal relationships from this observational study. As participants in the cohort develop AD, PD, FTD, or ALS, researchers can look for further associations between these diseases and various IDPs, and then plan further experiments to explain the relationships and identify risk/protective factors for diseases.
Ultimately, Miller said that similarly to any single nucleotide polymorphism in a genome-wide association study, no single measure in the U.K. Biobank will likely hold dramatic sway over disease risk. Rather, composites of multiple measures may emerge as robust risk factors for disease. To get a sense of what these composite measures could be, the researchers analyzed the imaging and other data to uncover “modes” of common characteristics among different people in the cohort. Each of the nine modes that emerged represented a cluster of IDPs and other health factors that significantly covaried among certain participants. In six of the modes, age was one such covariate. “Each of these [six] modes reflects a different aspect of the aging process,” Miller said. This underscores just how heterogeneous the aging process is, she added. That the researchers pulled out nine modes is a testament to the large size and depth of the data set. A similar analysis of data from the Human Connectome Project, which correlated brain imaging with 280 lifestyle and demographic variables in 461 healthy adults, only uncovered one mode (see Smith et al., 2015).
The modes brought out some curious correlations. For example, mode 7 linked low bone density and poor cognitive scores to multiple brain structure and diffusion MRI measures, including total brain volume. These relationships evoke previous findings linking weakening bones to cognitive decline and AD (see Yaffe et al., 1999; Tan et al., 2005). In mode 9, lifestyle factors such as eating more cheese or spending less time outdoors in the summer or winter correlated with cognitive measures such as fluid intelligence, as well as with a bevy of functional and structural MRI measures. Assigning meaning to these associations is tricky, Miller said, as some factors may simply be a proxy for socioeconomic status or education level. For example, people who eat more cheese may have a higher economic status, while spending time outdoors may reflect occupations that include heavy physical labor, which was also negatively associated with cognitive and imaging measures in this mode.
The researchers will monitor incident health outcomes, then see how those relate to the baseline brain imaging and non-imaging measures. While researchers will have indefinite access to participants’ medical records through the National Health Service, most participants will not be required to undergo regularly scheduled checkups or tests beyond the baseline measures. Rather, the occurrence of future health problems (such as dementia) will only be recorded when patients come to the doctor on their own volition, Miller said. Based on the demographics of the Biobank participants, the researchers estimated that by the time imaging is completed in 2022, around 1,800 people in the imaging cohort will show symptoms of AD, and 1,200 will develop PD. Five years later, those numbers are projected to rise to 6,000 and 2,800 for AD and PD, respectively, while 50-100 may have motor neuron disease. The large size of the cohort will facilitate discovery of presymptomatic markers for these diseases, particularly for rare ones such as ALS, Miller believes. The Biobank plans to scan 10,000 participants at least one more time, but Miller said researchers have yet to decide when to initiate this longitudinal substudy component.
U.K. Biobank may be the largest brain-imaging game in town, but other European studies play in the same league. The German National Cohort (GNC) has imaged the brains of 10,000 of its target of 30,000 participants, according to Fabian Bamberg of the University of Tuebingen, who heads the MRI portion of that study (see Bamberg et al., 2015). In addition to collection of health data, whole-body and cardiac imaging, the GNC’s brain-imaging protocol includes MRI measurements similar to the U.K. Biobank’s, with the exclusion of diffusion MRI and task-related functional MRI. Bamberg told Alzforum that researchers from the two projects are working together closely to harmonize the measurements they have in common. Similar in scope to the GNC study, the Rhineland Study is in the process of imaging 30,000 participants from the Bonn area, aged 30 and over, with plans to track their health outcomes for the remainder of their lifespans (see Breteler et al., 2014). The Maastricht Study, which so far has run more than 5,000 of its projected 10,000 participants through the MRI scanner, focuses specifically on factors associated with Type 2 diabetes and cardiovascular disease (see Schram et al., 2014). Authors of the U.K. Biobank study hope to synchronize their data with those from these other imaging studies in the future.
“U.K. Biobank will be a tremendous resource for studies on neurodegeneration, both for data discovery or for replication,” wrote Betty Tijms and Pieter-Jelle Visser of VU Medical Center in Amsterdam in a joint comment to Alzforum. “Because data collection is not focused on a single disease, it will be possible to find common mechanisms across different brain disorders.” However, they also note the shortcomings of the data set in terms of neurodegenerative disease research, such as the lack of PET scans and absence of participants older than 80.
While Miller considers U.K. Biobank’s broad scope a strength, other researchers criticized the £40 million price tag and its lack of focus on specific diseases. “This is a lot of normative data but no findings,” commented John Hardy of University College London. “Obtaining longitudinal data, especially in the context of PET imaging of amyloid and tau, would really make this a valuable resource.”
While the inclusion of PET imaging and/or CSF biomarker data in a cohort of this size would be a boon to neurodegeneration research, Miller explained that MRI is noninvasive and captures a broad range of information about the brain’s structure, function, and connectivity, while PET scans are expensive, trickier to execute, and expose participants to more ionizing radiation than X-rays. Bamberg gave similar reasons for excluding PET scans from the GNC study. Nick Fox of University College London agreed that PET scans or CSF biomarker studies would have added cost. “Essentially the tradeoff for U.K. Biobank is numbers versus depth—they have the power of very large numbers but that means that expensive, invasive, or lengthy assessments may not be justifiable,” Fox wrote to Alzforum.
Smaller imaging studies that do collect PET scans and CSF biomarkers are focused specifically on neurodegenerative disease, such as the Alzheimer’s Neuroimaging Initiative (ADNI), the Australian Imaging, Biomarker, and Lifestyle study, the Harvard Aging Brain Study, or the Mayo Clinic Study of Aging. Michael Weiner of the University of California, San Francisco, who heads ADNI, told Alzforum that despite its lack of AD-specific measures, the broad-based U.K. Biobank study is exciting for the neurodegenerative disease field.
However, he added that with large data sets come sizable hurdles in data interpretation. “In my view, one of the biggest challenges to medical science is to find more robust ways to reduce false positive and false negative results from analysis of large data sets,” Weiner wrote. “Nevertheless, this recent report from the U.K. Biobank is a ‘tour de force’ and all of us in the field look forward to seeing more exciting discoveries coming from this project.”—Jessica Shugart
References
Paper Citations
- Pirpamer L, Hofer E, Gesierich B, De Guio F, Freudenberger P, Seiler S, Duering M, Jouvent E, Duchesnay E, Dichgans M, Ropele S, Schmidt R. Determinants of iron accumulation in the normal aging brain. Neurobiol Aging. 2016 Jul;43:149-55. Epub 2016 Apr 13 PubMed.
- Smith SM, Nichols TE, Vidaurre D, Winkler AM, Behrens TE, Glasser MF, Ugurbil K, Barch DM, Van Essen DC, Miller KL. A positive-negative mode of population covariation links brain connectivity, demographics and behavior. Nat Neurosci. 2015 Nov;18(11):1565-7. Epub 2015 Sep 28 PubMed.
- Yaffe K, Browner W, Cauley J, Launer L, Harris T. Association between bone mineral density and cognitive decline in older women. J Am Geriatr Soc. 1999 Oct;47(10):1176-82. PubMed.
- Tan ZS, Seshadri S, Beiser A, Zhang Y, Felson D, Hannan MT, Au R, Wolf PA, Kiel DP. Bone mineral density and the risk of Alzheimer disease. Arch Neurol. 2005 Jan;62(1):107-11. PubMed.
- Bamberg F, Kauczor HU, Weckbach S, Schlett CL, Forsting M, Ladd SC, Greiser KH, Weber MA, Schulz-Menger J, Niendorf T, Pischon T, Caspers S, Amunts K, Berger K, Bülow R, Hosten N, Hegenscheid K, Kröncke T, Linseisen J, Günther M, Hirsch JG, Köhn A, Hendel T, Wichmann HE, Schmidt B, Jöckel KH, Hoffmann W, Kaaks R, Reiser MF, Völzke H, German National Cohort MRI Study Investigators. Whole-Body MR Imaging in the German National Cohort: Rationale, Design, and Technical Background. Radiology. 2015 Oct;277(1):206-20. Epub 2015 May 19 PubMed.
- Schram MT, Sep SJ, van der Kallen CJ, Dagnelie PC, Koster A, Schaper N, Henry RM, Stehouwer CD. The Maastricht Study: an extensive phenotyping study on determinants of type 2 diabetes, its complications and its comorbidities. Eur J Epidemiol. 2014 Jun;29(6):439-51. Epub 2014 Apr 23 PubMed.
External Citations
Further Reading
Papers
- Pilling LC, Atkins JL, Bowman K, Jones SE, Tyrrell J, Beaumont RN, Ruth KS, Tuke MA, Yaghootkar H, Wood AR, Freathy RM, Murray A, Weedon MN, Xue L, Lunetta K, Murabito JM, Harries LW, Robine JM, Brayne C, Kuchel GA, Ferrucci L, Frayling TM, Melzer D. Human longevity is influenced by many genetic variants: evidence from 75,000 UK Biobank participants. Aging (Albany NY). 2016 Mar;8(3):547-60. PubMed.
Primary Papers
- Miller KL, Alfaro-Almagro F, Bangerter NK, Thomas DL, Yacoub E, Xu J, Bartsch AJ, Jbabdi S, Sotiropoulos SN, Andersson JL, Griffanti L, Douaud G, Okell TW, Weale P, Dragonu I, Garratt S, Hudson S, Collins R, Jenkinson M, Matthews PM, Smith SM. Multimodal population brain imaging in the UK Biobank prospective epidemiological study. Nat Neurosci. 2016 Sep 19; PubMed.
Annotate
To make an annotation you must Login or Register.

Comments
San Francisco Veterans Affairs Medical Center
The U.K. Biobank is a fantastic project, and I congratulate the investigators and staff involved with designing, collecting, and analyzing its data. Considering that this is just the first step, the results are impressive and demonstrate the feasibility and statistical power of this hugely ambitious effort.
There are clearly many differences between ADNI and the U.K. Biobank. ADNI’s goal is to validate biomarkers for Alzheimer’s disease clinical trials, while the U.K. Biobank has much broader goals. ADNI includes amyloid tau PET imaging as well as analysis of cerebrospinal fluid obtained by lumbar punctures. The present paper from the U.K. Biobank project currently does not include amyloid or tau PET imaging on a large scale, but I believe there are plans to include these modalities in the future on a considerable number of subjects.
From the onset, we in ADNI recognized that the subject population in ADNI is not representative of the “general population”; in other words, ADNI is not an “epidemiologically sampled study.” The Olmsted County Study of Aging is a good example of a study, which uses MRI and PET imaging, that aims to represent the population living near the Mayo Clinic in Rochester, Minnesota. In contrast, ADNI is designed to represent a typical AD clinical trial in the United States, and its subjects are more educated and more Caucasian than the general population. For ADNI we rule out people with cerebrovascular disease or cognitive impairment caused by disorders other than AD, because AD clinical trials typically used these same exclusion criteria. Therefore, investigators must be careful not to overgeneralize ADNI findings when interpreting its results.
The relationships between age, cognition, brain volumes, brain amyloid, etc. in ADNI may or may not reflect that of the general population. One big advantage of the U.K. Biobank study is that its investigators have access to the large amount of health data that is collected by the U.K. universal health care system. In ADNI we lack access to such data.
Of course caveats apply to all research studies, especially those with lots of intensive measurements such as MRI. Even the U.K. Biobank can only study those subjects who are willing to enter into the study. People who are homeless, or who have severe diseases, or wish to avoid participating in research will most likely not be in the U.K. Biobank. It certainly will be very interesting to compare the results of the U.K. Biobank with those of ADNI and the Olmsted County Study of Aging.
Having been involved closely with ADNI for more than 12 years, I can share one other concern about these very large studies. It is a general concern about the results coming out of “big data” studies in general. The most robust scientific findings come when a carefully formulated a priori hypothesis is being tested. In fact, in the U.K. Biobank paper, a number of a priori hypotheses are tested, and that is terrific. Sometimes the most exciting discoveries occur outside of the process of a priori hypothesis testing, but we all must be very careful about false positive results emerging because of insufficient correction for multiple comparisons. At the same time, sometimes real results are obscured because corrections for multiplicity prevent assignment of statistical significance to them.
This general concern applies to many large studies, including ADNI and the U.K. Biobank. In my view, a key challenge to medical science is to find more robust ways to reduce false-positive and false-negative results from analysis of large data sets.
Nevertheless, this recent report from the U.K. Biobank is a “tour de force,” and all of us in the field look forward to seeing more exciting discoveries coming from this project.
Maastricht University; VU University Medical Centre
Amsterdam UMC, loc. VUmc
The U.K. Biobank will be a tremendous resource for studies on neurodegeneration, both for data discovery and for replication. In addition, it will be a useful reference set for single-subject imaging markers. We have not used the data set ourselves, but many of our colleagues either have already obtained data or plan to request data. Because data collection is not focused on a single disease, it will be possible to find common mechanisms across different brain disorders.
A limitation of the current data set is the lack of molecular brain markers, for example in cerebrospinal fluid or from PET imaging, but this type of data will possibly be collected in the future. The inclusion of subjects above 80 would also have been useful. In such a huge data set correlations may not always be meaningful. For example, the analysis in table 9 might provide an incentive to eat more cheese and less yogurt. The huge data set urges for the development of new statistical techniques for data mining. To avoid selective reporting of results, one could ask data requesters to submit a protocol specifying the analysis, which later can be linked to a publication.
Make a Comment
To make a comment you must login or register.