NEALS: Desperately Seeking ALS Biomarkers
Quick Links
28 October 2011. Researchers studying amyotrophic lateral sclerosis (ALS) have plenty of drug ideas to try, but a great problem in figuring out if they work or not. Right now there are no good tests to detect the disease or measure its progression. “This field has been screaming out for biomarkers,” said Robert Miller of the California Pacific Medical Center in San Francisco, who leads the Western ALS Consortium and is a member of the Northeast ALS Consortium (NEALS) (see Part 1 and Part 2). A coalition of more than 140 ALS investigators at 100 sites, NEALS has been hunting for good biomarkers for the better part of a decade. It is running projects on fluid proteins and metabolites, as well as new tests to quantify muscle function.
The creeping paralysis that marks ALS can first make itself known with a stumble, fingers skittering across the keyboard, or slurred words. Of course, many diseases could cause these symptoms; doctors must eliminate all other possibilities even as ALS spreads from one body part to the next. It can take a year just to diagnose ALS. This means that people starting clinical trials are already well into the process of spinal cord neurodegeneration—when it may be too late for drugs to make a big difference.
Another problem for trials is that scientists have no good way to quantify disease progression, and thus no obvious method to determine whether a drug is slowing it down. The muscle strength and breathing tests used in many trials often yield variable results. Even survival is not entirely reliable as a measure of a drug’s effectiveness; How long people with ALS live depends on many factors, including whether they receive supportive therapy such as a tracheostomy (see ARF related news story on Gordon et al., 2009). Plus, using death as an endpoint requires large, lengthy trials. Neither researchers nor people with ALS want to wait around for this result.
In the absence of validated biomarkers for ALS, NEALS currently uses a combination of outcome measures, including lung capacity, muscle strength, and the ALS Functional Rating Scale, which scores a person based on day-to-day activities such as getting dressed. NEALS also uses motor unit number estimation (MUNE), a tool to assess muscle function in an axonal-loss disease. MUNE measures the number of functioning motor units, which include the motor neuron and the muscle it innervates. Electrophysiologists calculate MUNE based on nerve responses, recorded with surface electrodes, to stimuli. NEALS co-founder Jeremy Shefner of the State University of New York (SUNY) Upstate Medical University in Syracuse is evaluating and improving MUNE for the consortium’s studies.
NEALS has applied MUNE in some studies, such as a trial of celecoxib (Shefner et al., 2007), but previous results were wildly variable and fraught with artifacts (Shefner, 2009). In a new study of MUNE methods, published in July, Shefner and colleagues were able to obtain more stable results by measuring activity at multiple points along a nerve (Shefner et al., 2011). People with ALS declined by an average of 9 percent on MUNE each month. This rapid drop makes MUNE a powerful tool to use in ALS Phase 2 trials, said Merit Cudkowicz, Shefner’s NEALS co-founder at the Massachusetts General Hospital (MGH) in Boston.
NEALS is also investigating the value of two other measures of muscle health and strength in ALS trials. The group recently completed a trial to evaluate the ATLIS (Accurate Test of Limb Isometric Strength) device. Developed by Patricia Andres of MGH-East in Charlestown, Massachusetts, ATLIS consists of a chair mounted to a frame and looks a bit like a home gym. People sit in the chair, and the device measures the force they exert when flexing different muscle groups. In a study recently accepted for publication in Muscle & Nerve, Cudkowicz and colleagues report that the device was convenient and precise in experiments with 20 healthy people and 10 with ALS.
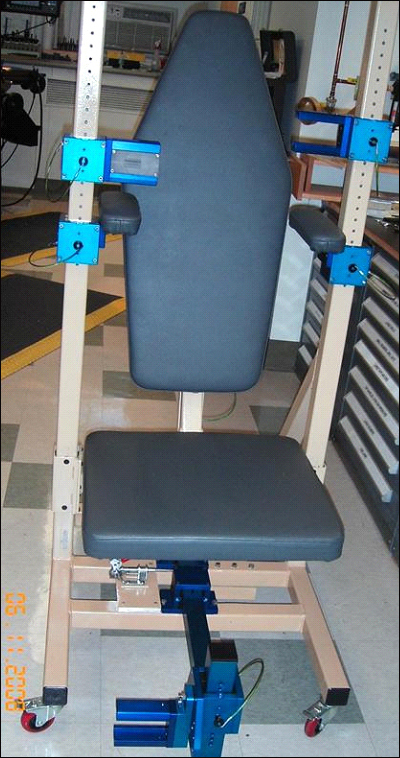
Using new devices such as the Accurate Test of Limb Strength (ATLIS), shown, may help clinicians better track the progression of ALS and any benefits from treatment. Image credit: NEALS
In addition, NEALS is collaborating with Seward Rutkove of the Beth Israel Deaconess Medical Center in Boston. Rutkove developed a handheld device that measures electrical impedance myography—in other words, how well electrical current travels through muscle (see ARF related news story; Rutkove, 2009).
Once a diagnosis of ALS is confirmed, MUNE, ATLIS, and the impedance device could turn out to be sufficiently sensitive measures to monitor disease progression during a clinical trial, Cudkowicz said. These measures are noninvasive and relatively portable. They are not specific for ALS pathology, but rather for motor unit degeneration; therefore, they could also serve research into other diseases. For example, researchers are using MUNE as an outcome measure in a trial for spinal muscular atrophy. The NEALS ATLIS study was sponsored in part by the Muscular Dystrophy Association based in Tucson, Arizona.
In the hunt for ALS-specific measures, Robert Bowser of the Barrow Neurological Institute in Phoenix, Arizona, has led NEALS’ molecular biomarker program for the better part of a decade. Much of the effort has been devoted to screening for protein markers, using mass spectrometry, in cerebrospinal fluid samples from ALS patients and healthy control donors. Promising candidates, Bowser said, include neurofilament, an important axonal component (Mendonça et al., 2011); inflammatory molecules such as complement component C3 (Goldknopf et al., 2006); and other proteins such as cystatin C (Wilson et al., 2010). (For an in-depth review on fluid biomarkers in neurodegenerative diseases, see Olsson et al., 2011.)
Now, researchers must verify those markers in large groups of people with and without the disease (Ryberg et al., 2010). Adding to the challenge, some diseases mimic ALS. Whereas ALS, by definition, affects both upper and lower motor neurons, it can be confused with similar conditions that affect only upper motor neurons, such as primary lateral sclerosis or hereditary spastic paraplegia, or only lower motor neurons, such as progressive muscular atrophy. In one NEALS study, the consortium collected samples from nearly 250 people with and without motor neuron disease to identify a marker that helps distinguish ALS from its mimics. Thus far, a combination of the phosphorylated neurofilament heavy chain and C3 best distinguishes ALS from these similar conditions. In a recent study, this pair correctly identified 87.3 percent of people with ALS among a population of 71 people with ALS, 40 healthy subjects, and 52 people with varied diseases including ALS mimics, multiple sclerosis, and other kinds of neurodegeneration (Ganesalingam et al., 2011).
The proteome is not the only hunting ground for biomarkers; small metabolites might flag ALS as well. NEALS is collaborating with Metabolon Inc. of Research Triangle Park, North Carolina, to seek metabolic markers of ALS in blood, and expects to publish results shortly, Bowser said (see ARF related news story).
At a 12 September 2011 joint workshop between NEALS and the ALS Association (ALSA), researchers met at Massachusetts General Hospital in Boston to discuss biomarker progress. “There is still work to be done,” noted Lucie Bruijn, ALSA chief scientist, in an e-mail to ARF. “As a community, we have done a good job of standardizing CSF and plasma collections, and are now working toward standardizing imaging techniques among centers.” Researchers are starting to use magnetic resonance imaging and positron emission tomography to examine degeneration of nerve fibers, for example, in the brains and spinal cords of people with ALS (see ARF related news story). Standardizing methods for CSF collection, biomarker measurements, and imaging has also been a crucial issue in Alzheimer’s research (see ARF related news story). It is a focus of the Alzheimer’s Disease Neuroimaging Initiative (ADNI), a longitudinal study in its seventh year of collecting and analyzing blood and urine samples, plus brain images, from 800-plus people with Alzheimer’s (see ARF related news story), as well as of a quality control initiative in which some 70 labs from around the world participate (see ARF related news story). Like ADNI and other studies, NEALS is starting to collect samples longitudinally to hunt for markers that track disease progression.
Many researchers expect there will not be one biomarker for ALS, but many. Several may work well in concert (see ARF related news story on Steinacker et al., 2011). In addition, Bowser suggested that researchers might want to use different biomarkers, depending on a drug’s action. If the medicine affects inflammation, then molecules in that pathway make the most sense; but an entirely different set of biomarkers could be preferable to monitor the efficacy of drugs protecting mitochondria, for example.—Amber Dance.
This is Part 3 of a four-part series. See also Part 1, Part 2, Part 4. Download a PDF of the entire series.
References
News Citations
- NEALS: Collaboration for a Cure to ALS
- NEALS: In ALS Trials, One Design Does Not Fit All
- Seeking a Reliable Yardstick for ALS Trials
- Metabolomics: Seeking Biomarkers for ALS, Parkinson’s, Huntington’s
- Portrait of a Motor Neuron Disease: Focus on Imaging for ALS
- Paris: Standardization a Hurdle for Spinal Fluid, Imaging Markers
- As ADNI Turns Four, $64 Million Data Start Rolling In
- Are Soluble APP Fragments Markers for ALS?
- NEALS: Sharing Among ALS Researchers and Participants
Paper Citations
- Gordon PH, Corcia P, Lacomblez L, Pochigaeva K, Abitbol JL, Cudkowicz M, Leigh PN, Meininger V. Defining survival as an outcome measure in amyotrophic lateral sclerosis. Arch Neurol. 2009 Jun;66(6):758-61. PubMed.
- Shefner JM, Cudkowicz ME, Zhang H, Schoenfeld D, Jillapalli D, . Revised statistical motor unit number estimation in the Celecoxib/ALS trial. Muscle Nerve. 2007 Feb;35(2):228-34. PubMed.
- Shefner JM. Statistical motor unit number estimation and ALS trials: the effect of motor unit instability. Suppl Clin Neurophysiol. 2009;60:135-41. PubMed.
- Shefner JM, Watson ML, Simionescu L, Caress JB, Burns TM, Maragakis NJ, Benatar M, David WS, Sharma KR, Rutkove SB. Multipoint incremental motor unit number estimation as an outcome measure in ALS. Neurology. 2011 Jul 19;77(3):235-41. PubMed.
- Rutkove S. Electrical impedance myography as a biomarker for ALS. Lancet Neurol. 2009 Mar;8(3):226; author reply 227. PubMed.
- Mendonça DM, Martins SC, Higashi R, Muscara MN, Neto VM, Chimelli L, Martinez AM. Neurofilament heavy subunit in cerebrospinal fluid: a biomarker of amyotrophic lateral sclerosis?. Amyotroph Lateral Scler. 2011 Mar;12(2):144-7. PubMed.
- Goldknopf IL, Sheta EA, Bryson J, Folsom B, Wilson C, Duty J, Yen AA, Appel SH. Complement C3c and related protein biomarkers in amyotrophic lateral sclerosis and Parkinson's disease. Biochem Biophys Res Commun. 2006 Apr 21;342(4):1034-9. PubMed.
- Wilson ME, Boumaza I, Lacomis D, Bowser R. Cystatin C: a candidate biomarker for amyotrophic lateral sclerosis. PLoS One. 2010;5(12):e15133. PubMed.
- Olsson B, Zetterberg H, Hampel H, Blennow K. Biomarker-based dissection of neurodegenerative diseases. Prog Neurobiol. 2011 Dec;95(4):520-34. PubMed.
- Ryberg H, An J, Darko S, Lustgarten JL, Jaffa M, Gopalakrishnan V, Lacomis D, Cudkowicz M, Bowser R. Discovery and verification of amyotrophic lateral sclerosis biomarkers by proteomics. Muscle Nerve. 2010 Jul;42(1):104-11. PubMed.
- Ganesalingam J, An J, Shaw CE, Shaw G, Lacomis D, Bowser R. Combination of neurofilament heavy chain and complement C3 as CSF biomarkers for ALS. J Neurochem. 2011 May;117(3):528-37. PubMed.
- Steinacker P, Fang L, Kuhle J, Petzold A, Tumani H, Ludolph AC, Otto M, Brettschneider J. Soluble beta-amyloid precursor protein is related to disease progression in amyotrophic lateral sclerosis. PLoS One. 2011;6(8):e23600. PubMed.
Other Citations
External Citations
Further Reading
Papers
- Levine TD, Bowser R, Hank N, Saperstein D. A pilot trial of memantine and riluzole in ALS: correlation to CSF biomarkers. Amyotroph Lateral Scler. 2010 Dec;11(6):514-9. PubMed.
- Ganesalingam J, Bowser R. The application of biomarkers in clinical trials for motor neuron disease. Biomark Med. 2010 Apr;4(2):281-97. PubMed.
- Cudkowicz ME, Swash M. CSF markers in amyotrophic lateral sclerosis: has the time come?. Neurology. 2010 Mar 23;74(12):949-50. PubMed.
- Bowser R, Lacomis D. Applying proteomics to the diagnosis and treatment of ALS and related diseases. Muscle Nerve. 2009 Nov;40(5):753-62. PubMed.
- Ryberg H, Bowser R. Protein biomarkers for amyotrophic lateral sclerosis. Expert Rev Proteomics. 2008 Apr;5(2):249-62. PubMed.
News
- Tracking MND Progression With New Model, New Marker
- Classifying Motor Neuron Disease With CSF Markers
- Portrait of a Motor Neuron Disease: Focus on Imaging for ALS
- Seeking a Reliable Yardstick for ALS Trials
- News Brief: Muscle-Monitoring Method Earns $1 Million ALS Prize
- Metabolomics: Seeking Biomarkers for ALS, Parkinson’s, Huntington’s
- Paris: Standardization a Hurdle for Spinal Fluid, Imaging Markers
- As ADNI Turns Four, $64 Million Data Start Rolling In
- Are Soluble APP Fragments Markers for ALS?
- NEALS: Collaboration for a Cure to ALS
- NEALS: In ALS Trials, One Design Does Not Fit All
Annotate
To make an annotation you must Login or Register.

Comments
No Available Comments
Make a Comment
To make a comment you must login or register.