Microglial Transcriptome Hints at Shortcomings of AD Model
Quick Links
With the realization that microglia play a key role in neurodegenerative conditions, scientists have profiled their transcription patterns to see how they change during disease. A study published in the January 16 Cell Reports pulls together and analyzes 18 such data sets. Led by David Hansen at Genentech, South San Francisco, California, the researchers confirmed the presence of a neurodegeneration-related phenotype in mouse models of Alzheimer’s disease and spotted the signature in human Alzheimer’s tissue. They also found a set of inflammation-related genes expressed in AD brains, but not in mouse models. The researchers hope to accelerate microglial research by sharing the compiled data in an interactive database online. Alzforum covered some of this work when first author Brad Friedman presented preliminary data at Microglia in the Brain, a 2016 Keystone symposium (Jul 2016 conference news).
- A new database integrates 18 data sets of microglial/myeloid expression profiles.
- A set of co-regulated genes associated with neurodegeneration in mice was spotted in AD brains.
- AD brains express a set of inflammation genes not seen in AD mice.
“These data sets are one of the most valuable things for the community,” said Shane Liddelow, New York University, who has already mined the database for evidence of cross talk between microglia and astrocytes. Oleg Butovsky, Brigham and Women’s Hospital in Boston, also considers it an important resource but wondered how the authors’ analyses of the compiled data advances understanding of microglia.
Scientists have uncovered several microglial phenotypes associated with neurodegeneration and AD in particular, including a microglial neurodegenerative phenotype (MGnD) and “DAMs,” a.k.a disease-associated microglia. How they relate to each other, and what they do in neurodegeneration, still needs clarification (Jun 2017 news; Sep 2017 news; Sep 2017 news).
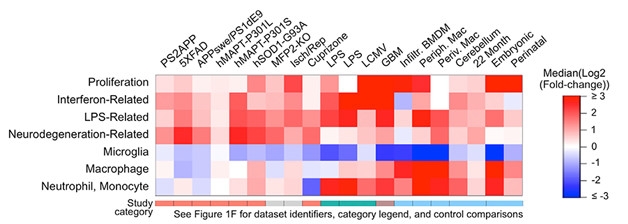
Multifaceted Personality. Different disease models and other conditions (columns) induce different myeloid cell expression modules in the brain (rows). Microglial gene expression is down or unchanged in most models of conditions, whereas a neurodegeneration-related signature is up in those respective models. (Courtesy of Friedman et al., Cell Reports, 2018.)
In search of a more global view of microglia’s multiple roles in disease, Friedman collected transcriptional data from models of ischemia, infectious disease, inflammation, cancer, demyelination, tauopathy, and Alzheimer’s disease. Most of the data came from published reports, but the authors also added new RNA-Seq data sets from the PS2APP model of AD, and from the P301L (Götz et al., 2001) and P301S tauopathy models.
To integrate all these data, Friedman used z-score normalization. For each study, he calculated how different disease conditions shifted expression of each gene from the average. Then he transformed the data into relative values that he could analyze together in one master gene expression matrix.
“One of the nice things about our paper is that it surveys a diverse set of microglial states. It’s trying to make sense of all those profiles, putting them all together,” said Hansen. Friedman added, “It’s also one of the first, maybe the first, to include a genome-wide expression data set of a tauopathy model.”
To identify common types of microglial activation in an unbiased way, Friedman selected 777 genes whose expression levels departed from the mean in the different models. He then used hierarchical clustering to group these genes into co-regulated modules. From 45 that emerged, the researchers focused on seven that were enriched for genes associated with cell proliferation, interferon signaling, responses to lipopolysaccharide (LPS), neurodegeneration, and functions of homeostatic microglia, macrophages, and neutrophils/monocytes (image above). The neurodegeneration profile proved similar to transcriptional changes seem in DAMs and MGnDs.
Reanalyzing single-cell data from the previous DAM study, Friedman and colleagues found 16 unique transcriptional signatures, suggesting discrete microglial states. Two of these signatures, one associated with DAMs and the other interferon-related, were more likely to show up in mouse models of AD than in controls. The interferon signature was roughly twice as abundant in 5xFAD mice as in controls.
The researchers also examined microglial expression in human AD tissue. They sequenced RNA from previously frozen samples of the fusiform gyrus from 33 Braak stage I-IV and 84 Braak stage III-VI cases from the Banner Research Institute tissue bank. The AD samples scored about 10 percent higher for the neurodegeneration module. The researchers also found an unexpected uptick in expression of the LPS module compared to mouse models, suggesting AD brains may have more inflammatory signaling and/or peripheral immune cell infiltration than do transgenic animals.
“It is common that when we look at animal models, they do not recapitulate well the glial aspects of disease,” said Liddelow.
Whether the postmortem findings truly reflect what happens in living AD brains remains to be seen, noted Michael Heneka, University of Bonn, Germany. “It is important to recognize that postmortem changes may be a strong confound, especially in AD, where microglia have been primed for decades.” Although the authors obtained tissue within five hours of death, microglial expression patterns can change in a matter of minutes (Jun 2017 news). “I think the most important next step is to get better expression data from human disease,” acknowledged Hansen.
Butovsky and Liddelow called for validation of the findings in situ. They would like to see how the microglia expression modules map to different locations in mouse and human brains, extending previous findings of disease-associated microglia. Indeed, Hansen is particularly interested in examining the distribution of cells expressing the interferon-related module. Heneka noted that it’s important to clarify the biological meaning of the transcriptional profiles, including the many untranslated transcripts. “Immune cells have a lot of bullets they never shoot,” he said.—Marina Chicurel
References
News Citations
- When a Microglia Is No Longer a Microglia
- Hot DAM: Specific Microglia Engulf Plaques
- ApoE and Trem2 Flip a Microglial Switch in Neurodegenerative Disease
- Microglial Kinase Promotes DAM, Blocks Lysosomal Aβ Digestion
- What Makes a Microglia? Tales from the Transcriptome
Research Models Citations
Paper Citations
- Götz J, Chen F, Barmettler R, Nitsch RM. Tau filament formation in transgenic mice expressing P301L tau. J Biol Chem. 2001 Jan 5;276(1):529-34. PubMed.
Other Citations
External Citations
Further Reading
Papers
- Yin Z, Raj D, Saiepour N, Van Dam D, Brouwer N, Holtman IR, Eggen BJ, Möller T, Tamm JA, Abdourahman A, Hol EM, Kamphuis W, Bayer TA, De Deyn PP, Boddeke E. Immune hyperreactivity of Aβ plaque-associated microglia in Alzheimer's disease. Neurobiol Aging. 2017 Jul;55:115-122. Epub 2017 Mar 27 PubMed.
Primary Papers
- Friedman BA, Srinivasan K, Ayalon G, Meilandt WJ, Lin H, Huntley MA, Cao Y, Lee SH, Haddick PC, Ngu H, Modrusan Z, Larson JL, Kaminker JS, van der Brug MP, Hansen DV. Diverse Brain Myeloid Expression Profiles Reveal Distinct Microglial Activation States and Aspects of Alzheimer's Disease Not Evident in Mouse Models. Cell Rep. 2018 Jan 16;22(3):832-847. PubMed.
Annotate
To make an annotation you must Login or Register.

Comments
Saba University School of Medicine
This study is a tour de force that provides significant insight into the complexity of gene expression profiles of AD and related mouse models, with a particular focus on microglial cells.
Several large studies have been performed recently analyzing gene expression changes in neurodegenerating brains, and almost all suffer from the same caveat. I will try to illustrate the problem with an example.
Imagine scientists interested in knowing the number of testicles per person in a human population. They are likely to conclude that statistically, on average, humans have one testicle per person. Although by itself a sound conclusion, from a biological point of view it is a silly one. We all know that males have two while females have none. Now if due to a disaster, the number of males in a population were to be reduced by half, these scientists could conclude that, based on total population data, the average number of testicles would fall to two-thirds per person. Obviously, on a per person level, still every man would have two and every woman would have none!
A similar problem exists in gene expression analyses performed on degenerating brains. In such a brain, there are some regions with up to 70 percent loss of neurons. Some past studies have concluded that in AD brains, APP gene expression levels are similar or lower compared to those of healthy brains. Far-reaching conclusions are formulated from such results in the context of production and clearance of APP and its derived peptides.
Such conclusions, however, are likely to be as silly as the two-thirds testicle per person conclusion. If just 30 percent of neurons show a somewhat decreased APP mRNA level, this probably indicates that on a per neuron level APP levels have been significantly increased in many different neuron types in such a brain. Also, other conclusions, like the level of gliosis and inflammation observed in AD brains, are likely to be significantly exaggerated when taking into account that some of the brain regions analyzed have lost a significant number of neurons.
No wonder little overlap is found between mouse models of AD and human AD. The mouse models do not show the significant amount of neuronal cell death and the gene expression patterns are significantly different.
Too many of the neurodegeneration-related changes in expression profiles from whole tissue samples are the result of altered cell-type composition and not due to transcriptional regulation of the genes involved. The field of neurodegeneration research has to make a significant effort to find experimental setup or analysis methods that capture gene expression in brains on a per cell type basis. The "whole tissue homogenate" approach produces too many misleading and erroneous results. Not really knowing if APP is under- or overexpressed in neurons of degenerating brains, or if microglia genes are overexpressed or downregulated on a per cell basis, is not acceptable anymore. A cell-type level of analysis should become the norm as soon as possible.
As a field, scientists need to come together and find some kind of consensus on how to deal with the cell type heterogeneity of the degenerating brain.
Make a Comment
To make a comment you must login or register.