From Garbage Disposal to Neuromodulation? Membrane Proteasomes Churn Out Stimulating Peptides
Quick Links
Long viewed as mere trash recyclers inside the cell, proteasomes—at least those in neurons—just got a shiny makeover. According to a paper in the March 13 Nature Structural and Molecular Biology, nearly half of neuronal 20S proteasomes reside not in the cytoplasm, but on the cell surface. From there, these neuronal membrane proteasomes (NMPs) devour intracellular proteins and spit peptides out into the extracellular space. Some of those peptides rapidly induce calcium influx, in part through activation of NMDA receptors on neurons, the researchers claim. Seth Margolis at Johns Hopkins University in Baltimore led the study, with a single graduate student, Kapil Ramachandran, conducting the experiments. The researchers have yet to identify the NMP’s substrates or products; however, if their finding holds up, the discovery of this novel proteasome will likely have wider implications for neuroscience, others said.
Alfred Goldberg of Harvard Medical School, whose work uncovered the function of the proteasome, was impressed. “This work suggests there’s a lot going in the cell membrane that we never dreamed of,” he said. Felix Schweizer of the University of California, Los Angeles, called the paper a tour de force. “Importantly, the authors have uncovered an intriguing and unexpected novel mechanism for communication by neurons. Since communication is crucial for the proper functioning of the brain, any new insight into neuronal communication does potentially bear on disease states. It is unexpected discoveries such as the ones made in this study that are enabled by fundamental research” (see full comment below).
Michael Ehlers of Biogen in Cambridge, Massachusetts, commented that the findings help explain previous observations that proteasome inhibition affects neuronal function with uncanny speed. “Ramachandran and Margolis provide evidence for a surprising and very different role of the proteasome in neurons,” he wrote. “It will be important to understand the molecular complex of the neuronal membrane proteasome in much more detail.”

Plasma Proteasome. Three potential models for how neuronal membrane proteasomes function within or at the plasma membrane. [Image courtesy of Ramachandran and Margolis, Nature Structural and Molecular Biology, 2017.]
The proteasome is the multi-subunit processor that degrades a majority of proteins in the cell. In the largest, 26S, version, ubiquitin-tagged proteins gain entry into the barrel-shaped, 20S, core after opening a regulatory cap (see Coux et al., 1996; Voges et al., 1999). The ubiquitin-proteasome system rids the cell of unwanted proteins, including some amyloidogenic ones implicated in neurodegenerative diseases. It is essential for many neuronal functions, including synaptic remodeling (see Ehlers, 2003).
Blocking the proteasome affects most processes on a timescale of hours to days. However, blocking the proteasome also dampens neurotransmission and alters the strength of the early phase of long-term potentiation within seconds to minutes. These rapid changes are hard to explain based on the complex’s effects on proteostasis (see Dong et al., 2008; Cai et al., 2010; Rinetti et al., 2010).
As a new graduate student in Margolis’s lab, Ramachandran noted a strong reduction in calcium signaling within seconds of blocking the proteasome. To investigate, he took a step back and asked where proteasomes reside in neurons. Using immunogold-electron microscopy, he probed hippocampal brain slices and primary hippocampal neurons for 20S core proteins, and was surprised to find about 40 percent of the 20S proteasomes smack in or near the plasma membrane. Antibodies for the 19S cap did not label the membrane, suggesting no 26S proteasome there. In contrast, HEK293 kidney cells had no membrane-associated 20S proteasomes.
Could these membrane-bound proteasomes somehow explain the rapid cellular responses to proteasome inhibitors? To test this, Ramachandran treated primary mouse cortical neurons with a cell-impermeable proteasome inhibitor called biotin-epoxomicin. This inhibitor is exquisitely specific for the proteasome and only binds to active ones, noted Goldberg. Within 10 to 30 seconds it strongly suppressed the amplitude of calcium transients induced by the GABA antagonist bicuculline. Biotin-epoxomicin also messed with the frequency of the transients, increasing it in half the neurons and decreasing it in a third.

Perky Peptides.
Neurons expressing the calcium indicator GCaMP light up when inhibitory transmission is erased with bicuculline (top). Blocking the neuronal membrane proteasome with biotin-epoxomicin prevents this response (bottom). [Image courtesy of Ramachandran and Margolis, Nature Structural and Molecular Biology, 2017.]
Ramachandran and Margolis reasoned that the NMPs might release peptides into the extracellular space, which then act as neuromodulators. Sure enough, through radioactive protein labeling experiments, Ramachandran found that cultured neurons churned out peptides into the medium and that proteasome inhibitors blocked this exodus. Isolating these peptides, he found they triggered calcium transients when added to primary mouse cortical neurons. Proteinase K abolished this effect. Also, peptides from neurons treated with proteasome inhibitors were unable to trigger calcium influx, suggesting only peptides coming out of the proteasome have this capability. Finally, using various inhibitors of ion channels and neurotransmitter receptors, the researchers determined that NMP peptides triggered the influx of calcium in part via NMDA receptors.
Margolis said this activity of NMP peptides may explain why the proteasome has been implicated in acute neuronal signaling. “Usually we think of signal amplification as happening from outside-in, but this is the opposite,” Margolis said. “Small changes inside the cell translate to massive changes outside.” Margolis added that it is possible the NMPs also work bidirectionally, processing proteins from the outside of the cell inwards, although that has yet to be formally tested.
What holds the 20S proteasome at the membrane? Ramachandran found that antibodies to proteasome subunits readily stained the surface of live neurons, and that the proteasome was sensitive to proteases added to the culture medium. These experiments indicated that at least some of the proteasome protrudes through to the extracellular side of the membrane. Biochemical experiments confirmed that the 20S proteasome clung tightly to membranes in neurons. Moreover, the multipass transmembrane glycoprotein GPM6A/B, expressed primarily in neurons, appears to anchor the NMPs to the membrane. Ramachandran thinks the NMPs, which are hydrophilic, might sit in a pore in the membrane formed by GPM6A/B (see image above).
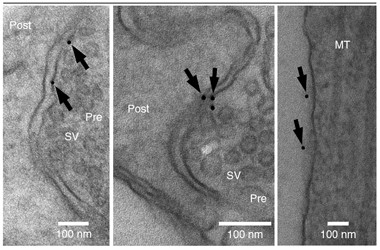
Reagent Gold.
Streptavidin-gold staining of neurons treated with biotin-epoxomicin suggest active proteasomes exist on synaptic membranes (arrows). [Image courtesy of Ramachandran and Margolis, Nature Structural and Molecular Biology, 2017.]
Why has this membrane proteasome gone unnoticed? Ramachandran said one contributing factor is scientists’ tendency to focus on the capped 26S proteasome rather than uncapped 20S cores. Goldberg added that while all cells contain uncapped 20S cores, no role for these “free agents” had been convincingly demonstrated before. Furthermore, when isolating proteasomes, researchers often throw away membrane fractions from the get-go, Goldberg said. Even with the high resolution of electron microscopy, it is difficult to confirm that a protein is actually associated with the membrane, not merely close by, Ramachandran added.
The findings raise many other questions. How are NMPs regulated? What are their substrates and products? Do NMPs process misfolded proteins implicated in neurodegenerative disease? Finally, do peptides released by NMPs contribute to the heightened neural activity observed during neurodegeneration? How NMPs relate to other degradation pathways in the cell, including the lysosomal and autophagic systems that tend to become overwhelmed during neurodegenerative disease, will be important to understand as well, Ramachandran and Margolis added.
Thomas Behnisch of Fudan University in Shanghai commented that the discovery of the NMP could be important for future and past neurodegenerative disease research. “Reduced proteasome activity has been shown to attenuate some forms of memory,” Behnisch wrote. “However, with the discovery of the transmembrane proteasome, some of the conclusions drawn from previous studies might need to be re-evaluated to take the new findings into account.”
Goldberg noted the proteasome’s essential role in generating antigenic peptides that are then loaded onto major histocompatibility complexes. Much as these peptides allow for constant immunosurveillance of the intracellular environment by lymphocytes, perhaps these NMP-generated peptides provide a continuous stream of information about the activity of synapses or dendrites, he suggested. Whether and how other cells, including neurons and glia, pick up on that information would have to be investigated, he said. Goldberg added that NMP-generated peptides might be externally loaded onto MHC I molecules on neurons or nearby cells. In addition to their classical role in presenting antigens to lymphocytes, MHC I molecules have also been implicated in neuronal development and plasticity and may protect motor neurons from degradation in people who have amyotrophic lateral sclerosis (see Huh et al., 2000; Mar 2016 news). Interestingly, the researchers found that NMPs included subunits specific to the immunoproteasome, which are found in antigen-presenting, hematopoietic cells and specialize in producing antigenic peptides.—Jessica Shugart
References
News Citations
Paper Citations
- Coux O, Tanaka K, Goldberg AL. Structure and functions of the 20S and 26S proteasomes. Annu Rev Biochem. 1996;65:801-47. PubMed.
- Voges D, Zwickl P, Baumeister W. The 26S proteasome: a molecular machine designed for controlled proteolysis. Annu Rev Biochem. 1999;68:1015-68. PubMed.
- Ehlers MD. Activity level controls postsynaptic composition and signaling via the ubiquitin-proteasome system. Nat Neurosci. 2003 Mar;6(3):231-42. PubMed.
- Cai F, Frey JU, Sanna PP, Behnisch T. Protein degradation by the proteasome is required for synaptic tagging and the heterosynaptic stabilization of hippocampal late-phase long-term potentiation. Neuroscience. 2010 Sep 15;169(4):1520-6. Epub 2010 Jun 22 PubMed.
- Rinetti GV, Schweizer FE. Ubiquitination acutely regulates presynaptic neurotransmitter release in mammalian neurons. J Neurosci. 2010 Mar 3;30(9):3157-66. PubMed.
- Huh GS, Boulanger LM, Du H, Riquelme PA, Brotz TM, Shatz CJ. Functional requirement for class I MHC in CNS development and plasticity. Science. 2000 Dec 15;290(5499):2155-9. PubMed.
Further Reading
Papers
- Bingol B, Schuman EM. Activity-dependent dynamics and sequestration of proteasomes in dendritic spines. Nature. 2006 Jun 29;441(7097):1144-8. PubMed.
Primary Papers
- Ramachandran KV, Margolis SS. A mammalian nervous-system-specific plasma membrane proteasome complex that modulates neuronal function. Nat Struct Mol Biol. 2017 Mar 13; PubMed.
Annotate
To make an annotation you must Login or Register.

Comments
UCLA
The work is amazing in scope and the diversity of methods and approaches that have been employed by the two authors and conducted experimentally only by the first author! A true tour de force. The work introduces a completely novel and unexpected potential for neurons to communicate with each other and with other cells. The discovery that in neurons a subset of proteasomes spans the plasma membrane opens the potential for the localized production of signaling peptides that are secreted via a non-vesicular mechanism. Besides the necessary confirmation of the results by other groups, the most important questions in my mind concern the nature of the peptides generated, what action(s) they have and on which cells, and whether neuronal activity could alter the amount or type of peptides—questions that the authors are raising in the discussion.
The work represents a fantastic example of fundamental research. Whether activity of the neuronal membrane proteasome (NMP) might be good or bad for neuronal health is not yet known. So it seems a bit premature to speculate on what this fundamental discovery means for diseases. The authors have uncovered an intriguing and unexpected novel mechanism for communication by neurons. Since communication is crucial for the proper functioning of the brain, any new insight into neuronal communication does potentially bear on diseases states. In a disease state, the NMP could be blocked by misfolded proteins either from the cytoplasm or from the extracellular side. Alternatively, disease-linked modulation of the NMP activity or the types of peptides produced could have deleterious effects on neurons or other cells. It is unexpected discoveries such as the ones made in this study that are enabled by fundamental research. Fundamental research leads to a better understanding of and unexpected insights into how the brain works. Such deeper understanding has many benefits, including novel insights into disease mechanisms.
The authors have substantiated their findings by testing them with a diverse set of techniques, each with different caveats. This softens the fact that many experiments were only done twice or three times, making traditional statistical claims questionable. I am much looking forward to an analysis of the peptides that are produced by the NMP. Knowing the identity of the peptides will go some way toward elucidating the function of the NMP. Unfortunately, the analysis of the action of the peptides is very preliminary and the calcium spikes observed in Fig 6 are not directly comparable to the sustained calcium increases in described in Fig 7. Maybe the processed peptides used in Fig 7 are distinct from the “fresh” peptides in Fig 6? Similar caution seems advised when considering whether NMDA receptors are involved in peptide action (Fig 7).
Fudan University
The authors discovered that some of the 20S proteasomes are localized by a glycoprotein sheath within the membranes of neurons; reaching out into the extracellular space. The specific transmembrane location of these 20S proteasomes allows them to degrade intracellular proteins by proteolysis and to secrete peptides into the interstitial space. The experiments show that the peptides of the proteolysed proteins regulate neuronal functions, such as synaptic transmissions and calcium signaling. The discovery of the transmembrane proteasome that secretes peptides with functional activity is, therefore, interesting for basic and medical researchers. The proteasome has been in focus in clinical research as a target for anti-cancer treatment and for Alzheimer’s and Parkinson’s diseases. Reduced activity of the proteasome has been shown to attenuate some forms of memory formation. However, with the discovery of the transmembrane proteasome, some of the conclusions drawn from previous studies might need to be re-evaluated to take the new findings into account. Thus, ongoing research has to show if the transmembrane proteasome represents a useful target to battle cellular processes of neurodegeneration.
Washington University
This is an interesting series of studies. First, I’ve always thought of the proteasome as a slow processor that degrades a wide variety of things in a cell. That it can act rapidly to modulate neuronal activity is interesting. But more intriguing to me is that this is another way for a cell to secrete proteins into the brain’s extracellular space, potentially in a synaptic-dependent manner. Many neurodegenerative diseases involve proteins found within the extracellular space, but in many cases we don’t know how or why they are secreted. For Aβ, we know how the peptide gets secreted and it accumulates in extracellular space, so that makes sense to some extent. But tau and α-synuclein are generally intracellular proteins that accumulate intracellularly. How those proteins are secreted into the extracellular space, and why they are there, remains largely unknown. (For that matter, why are many proteins in the extracellular space, such as GFAP?) A proteasome that links an intracellular compartment with the extracellular space could be a mechanism that eliminates these proteins from the cell. Additionally, Aβ, tau, and α-synuclein secretion are all regulated by neuronal activity. Perhaps this neuronal membrane proteasome is playing a role in synaptic-dependent secretion. Much of this above is speculation, but it is certainly fun to think about how this new proteasome function could impact neurodegenerative diseases.
University of Arkansas for Medical Sciences
Interesting and perhaps related: It has been known for some time that extracellular UPS components are present and functional (Sixt et al, 2008), including in cerebrospinal fluid (Mueller et al., 2012). This may reflect favorably on Model 2 proposed by Ramachandran and Margolis.
References:
Sixt SU, Beiderlinden M, Jennissen HP, Peters J. Extracellular proteasome in the human alveolar space: a new housekeeping enzyme?. Am J Physiol Lung Cell Mol Physiol. 2007 May;292(5):L1280-8. Epub 2007 Jan 12 PubMed.
Mueller O, Anlasik T, Wiedemann J, Thomassen J, Wohlschlaeger J, Hagel V, Keyvani K, Schwieger I, Dahlmann B, Sure U, Sixt SU. Circulating Extracellular Proteasome in the Cerebrospinal Fluid: A Study on Concentration and Proteolytic Activity. J Mol Neurosci. 2011 Sep 1; PubMed.
Make a Comment
To make a comment you must login or register.