Crystal Structure Suggests Nicastrin Binds γ-Secretase Substrates
Quick Links
By solving the crystal structure of nicastrin—one of the quartet of proteins that form γ-secretase—scientists reveal more detail than ever before about this protease complex. Yigong Shi and colleagues at Tsinghua University in Beijing zoomed in to atomic resolution to see the intricate details of nicastrin. The results offer new evidence that the subunit captures substrates that are then chopped up by the catalytic component, presenilin. With these new structural details in hand, the authors propose a mechanism for how the protease works. The results appeared September 16 in the Proceedings of the National Academy of Sciences.
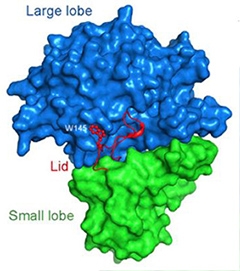
A Bi-lobed structure.
Nicastrin boasts a large and a small lobe. A loop (red) from the small lobe covers a cleft in the large one, like a lid. [Courtesy of Xie et al., PNAS.]
Recent evidence strengthens the argument that nicastrin binds substrates for γ-secretase (see May 2012 news story and Shah et al., 2005), but that idea still stirs debate. A clearer view of nicastrin’s structure could help settle the matter. Shi’s group previously used cryo-electron microscopy to visualize human γ-secretase at 4.5 Å (see Lu et al. 2014). However, components of this enzyme complex have proven difficult to crystallize and study at atomic-level detail. Instead, the authors turned to the slime mold Dictyostelium purpureum, a single-celled eukaryote with a γ-secretase that contains four subunits homologous to the human version. This amoeba version of the enzyme can process human APP to Aβ40 or Aβ42. Using X-ray crystallography, the researchers now resolve structural details down to 1.95 Å.
First authors Tian Xie and Chuangye Yan report that nicastrin from D. purpureum has one small and one large lobe, each consisting of numerous α-helices and β-strands. At the interface, hydrophobic phenyl rings from the large lobe protrude into a “greasy” pocket formed by lipophilic side chains from the small one, like a bolt in a receiver. This hinge likely allows the two subunits to pivot relative to one another, wrote the authors. The group also discovered that a loop extending from the small lobe covers the putative substrate-binding site on the large one, much like a lid (see image above). Prior low-resolution techniques overlooked this feature.
To help elucidate the function of nicastrin, the authors searched for similar structures in the Protein Data Bank. The closest was a bacterial aminopeptidase with homologous large and small lobes. The large one contains two zinc ions essential for the catalytic activity of this enzyme. The homologous site on nicastrin lacks amino acids that can bind zinc. This likely explains why nicastrin has no protease activity of its own, the authors wrote. However, they suggest that the protein could still bind substrates, and potentially deliver them to presenilin.
In contrast to nicastrin's lobes, those in the aminopeptidase keep the “lid” away from the active site, leaving it permanently open. The authors predict that the lid in nicastrin might have a regulatory role. “Our structural analysis of nicastrin provides a tantalizing clue about its function,” they write. They speculate that in an inactive state, the lid covers the substrate-binding pocket. When the two lobes pivot around the hydrophobic link, the lid opens, allowing nicastrin to bind substrates (see image below).

Open Sesame. At left, the “lid” (red) covers the substrate-binding site (green) on nicastrin. When the large lobe rotates around the hydrophobic pivot (blue), the lid opens, allowing substrate (yellow) to bind. [Courtesy of Xie et al., PNAS.]
“This new study fuels speculation that nicastrin may indeed serve as a receptor of γ-secretase substrates,” wrote David Bolduc and Michael Wolfe, Brigham and Women’s Hospital, Boston, in an accompanying commentary published online September 29. More work will be required to prove that hypothesis, they wrote. “Gaining a detailed mechanistic understanding of how γ-secretase processes APP—and its other substrates, such as Notch receptors—is critically important for biology and medicine.” Scientists should obtain high-resolution structures of the other components of γ-secretase as well, especially the catalytic subunit presenilin, they added, which is the focus of modulators that are being tested as therapeutics (see Jul 2014 news story). More details will illuminate ways to target this enzyme for Alzheimer’s disease, they wrote.
“This nicastrin structural model clearly supports previous reports that γ-secretase accesses its substrates through nicastrin,” wrote Xulun Zhang, University of Chicago, to Alzforum in an email (see full comment below). He was particularly interested in the lid, calling it unexpected and wondering if it serves only as an on/off switch, or if it could play additional roles in recruiting substrates.—Gwyneth Dickey Zakaib
References
News Citations
Paper Citations
- Shah S, Lee SF, Tabuchi K, Hao YH, Yu C, LaPlant Q, Ball H, Dann CE, Südhof T, Yu G. Nicastrin functions as a gamma-secretase-substrate receptor. Cell. 2005 Aug 12;122(3):435-47. PubMed.
- Lu P, Bai XC, Ma D, Xie T, Yan C, Sun L, Yang G, Zhao Y, Zhou R, Scheres SH, Shi Y. Three-dimensional structure of human γ-secretase. Nature. 2014 Aug 14;512(7513):166-70. Epub 2014 Jun 29 PubMed.
External Citations
Further Reading
Papers
- Tomita T. Molecular mechanism of intramembrane proteolysis by γ-secretase. J Biochem. 2014 Oct;156(4):195-201. Epub 2014 Aug 9 PubMed.
- Delay C, Dorval V, Fok A, Grenier-Boley B, Lambert JC, Hsiung GY, Hébert SS. MicroRNAs targeting Nicastrin regulate Aβ production and are affected by target site polymorphisms. Front Mol Neurosci. 2014;7:67. Epub 2014 Jul 18 PubMed.
- Holmes O, Paturi S, Selkoe DJ, Wolfe MS. Pen-2 is essential for γ-secretase complex stability and trafficking but partially dispensable for endoproteolysis. Biochemistry. 2014 Jul 15;53(27):4393-406. Epub 2014 Jun 30 PubMed.
- Lee SH, Sharma M, Südhof TC, Shen J. Synaptic function of nicastrin in hippocampal neurons. Proc Natl Acad Sci U S A. 2014 Jun 17;111(24):8973-8. Epub 2014 Jun 2 PubMed.
Primary Papers
- Xie T, Yan C, Zhou R, Zhao Y, Sun L, Yang G, Lu P, Ma D, Shi Y. Crystal structure of the γ-secretase component nicastrin. Proc Natl Acad Sci U S A. 2014 Sep 16;111(37):13349-54. Epub 2014 Sep 2 PubMed.
- Bolduc DM, Wolfe MS. Structure of nicastrin unveils secrets of γ-secretase. Proc Natl Acad Sci U S A. 2014 Oct 14;111(41):14643-4. Epub 2014 Sep 29 PubMed.
Annotate
To make an annotation you must Login or Register.

Comments
University of Chicago
In this study, Xie et al. tell us a structural tale of two lobes with the atomic resolution structure of nicastrin, the largest component of γ-secretase. Here, the ectodomain (ECD) of nicastrin from Dictyostelium purpureum, whose γ-secretase can faithfully process human APP (McMains et al., 2010), was expressed, purified, and crystalized. The structure of the ECD crystal was then solved at 1.95Å. Xie et al. showed that nicastrin contains one large lobe and one small, with each containing smaller structural domains. These two lobes are held together by multiple hydrogen bonds and Van der Waals interactions. A pocket surrounded by hydrophilic residues is located in the large lobe, which is speculated to be the binding site for γ-secretase substrates. A short loop extending out from the small lobe covers this pocket, seemingly a lid for this potential substrate binding site.
Based on the above information, Xie et al. also refined the 4.5 Å human nicastrin structural model, which was reported by the same research group earlier this year through cryo-electron-microscopy single-particle analysis (Lu et al., 2014). They showed that human and Dictyostelium purpureum nicastrins share many highly conserved structures, including the bilobed overall structure, similar H-bonds and Van der Waals interactions between the two lobes, a speculated substrate binding pocket in the large lobe and a lid covering this pocket. In addition, they redefined the relative positions of other components in γ-secretase.
In this structure model, since the lid blocks the potential substrate-binding pocket, they speculated that the model represents an inactive form of nicastrin. In order for nicastrin to capture substrate, Xie et al. propose that the lid could be removed through the relative rotation of the large and small lobes.
This nicastrin structural model clearly supports previous reports that γ-secretase accesses its substrates through nicastrin (Shah et al., 2005; Dries et al., 2009; Zhang et al., 2012). Knowing the positions of those residues surrounding the pocket in the large lobe will help to design experiments to further confirm the function of this pocket. The unexpected lid structure is another interesting discovery. Does the lid serve simply as a switch to convert nicastrin between inactive and active conformation? Or, does it also play other roles in recruiting substrates? After all, there is a long list of substrates trying to access γ-secretase through this binding site. Understanding how the lid functions will help elucidate how nicastrin recruits substrates. As another potential therapeutic target in addition to presenilin, the accurate structural information for nicastrin will certainly contribute to new ideas for modulating γ-secretase activity. This study, together with the EM study on γ-secretase complex, undoubtedly advanced our knowledge of the structure of this important enzyme.
References:
McMains VC, Myre M, Kreppel L, Kimmel AR. Dictyostelium possesses highly diverged presenilin/gamma-secretase that regulates growth and cell-fate specification and can accurately process human APP: a system for functional studies of the presenilin/gamma-secretase complex. Dis Model Mech. 2010 Sep-Oct;3(9-10):581-94. PubMed.
Lu P, Bai XC, Ma D, Xie T, Yan C, Sun L, Yang G, Zhao Y, Zhou R, Scheres SH, Shi Y. Three-dimensional structure of human γ-secretase. Nature. 2014 Aug 14;512(7513):166-70. Epub 2014 Jun 29 PubMed.
Shah S, Lee SF, Tabuchi K, Hao YH, Yu C, LaPlant Q, Ball H, Dann CE, Südhof T, Yu G. Nicastrin functions as a gamma-secretase-substrate receptor. Cell. 2005 Aug 12;122(3):435-47. PubMed.
Dries DR, Shah S, Han YH, Yu C, Yu S, Shearman MS, Yu G. Glu-333 of nicastrin directly participates in gamma-secretase activity. J Biol Chem. 2009 Oct 23;284(43):29714-24. PubMed.
Zhang X, Hoey RJ, Lin G, Koide A, Leung B, Ahn K, Dolios G, Paduch M, Ikeuchi T, Wang R, Li YM, Koide S, Sisodia SS. Identification of a tetratricopeptide repeat-like domain in the nicastrin subunit of γ-secretase using synthetic antibodies. Proc Natl Acad Sci U S A. 2012 May 29;109(22):8534-9. PubMed.
Make a Comment
To make a comment you must login or register.